The IntFOLD Server Results
Please cite the following papers:McGuffin, L.J., Atkins, J., Salehe, B.R., Shuid, A.N. & Roche, D.B. (2015) IntFOLD: an integrated server for modelling protein structures and functions from amino acid sequences. Nucleic Acids Research, 43, W169-73. PubMed
Roche, D. B., Buenavista, M. T., Tetchner, S. J. & McGuffin, L. J. (2011) The IntFOLD server: an integrated web resource for protein fold recognition, 3D model quality assessment, intrinsic disorder prediction, domain prediction and ligand binding site prediction. Nucleic Acids Res., 39, W171-6. PubMed
Links to graphical output:
- Top 5 3D models
- Disorder prediction
- Domain boundary prediction
- Binding site prediction
- Full model quality assessment results
Download machine readable results in CASP format:
- TS (Tertiary Structure Prediction)
- DR (Disorder Prediction)
- DP (Domain Prediction)
- FN (Binding Site Prediction)
- QA (Model Quality Prediction)
Results will be available for 21 days after job completion (subject to server capacity)
| Top 5 3D models for T0585 | Help | ||||
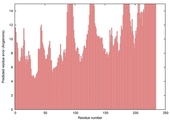
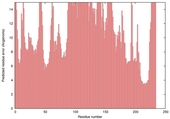
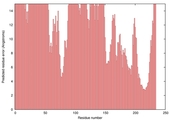
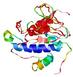
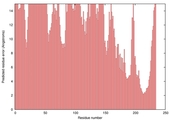
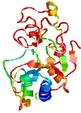
| Model name (PDBsum links for templates used) | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | Model coloured by local quality (click images to view models, local errors and target coverage interactively) |
| nFOLD4_multi _HHsearch_TS1 1jwqA 1xovA |
MEDIUM: 1.208E-2 |
0.2510 |
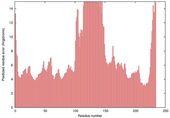 |
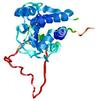 |
| nFOLD4_1jwqA _COMA_TS1 |
MEDIUM: 1.395E-2 |
0.2429 |
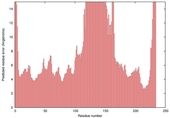 |
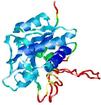 |
| nFOLD4_1jwqa _spk2_TS1 |
MEDIUM: 1.493E-2 |
0.2391 |
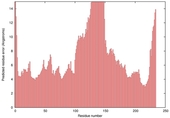 |
 |
| nFOLD4_1jwqa _sp3_TS1 |
MEDIUM: 1.493E-2 |
0.2391 |
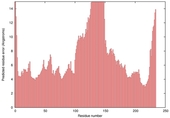 |
 |
| nFOLD4_1xovA _HHsearch_TS2 |
MEDIUM: 1.637E-2 |
0.2339 |
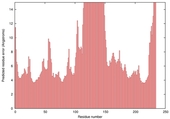 |
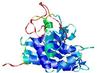 |
McGuffin, L. J. (2008) Intrinsic disorder prediction from the analysis of multiple protein fold recognition models. Bioinformatics, 24, 1789-1804. PubMed
| Disorder prediction for T0585 | Help | |||
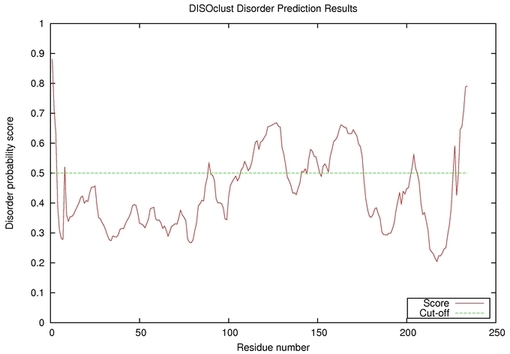
| |||
| Click image to download plot in PostScript format. |
| Domain boundary prediction for T0585 | Help | |||
| |||
| Click image to download model or view prediction using Jmol. The model above is coloured according to the predicted domains. |
Roche, D. B., Tetchner, S. J. & McGuffin, L. J. (2011) FunFOLD: an improved automated method for the prediction of ligand binding residues using 3D models of proteins. BMC Bioinformatics, 12, 160. PubMed
| Ligand binding residues prediction for T0585 | Help | |||
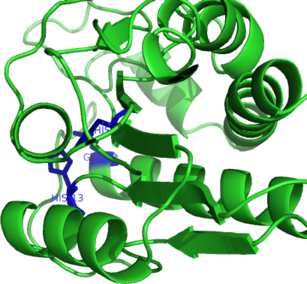
| |||
| Click image to download model or view prediction using Jmol. Predicted ligand binding residues are shown as blue sticks in the image above. |
McGuffin, L. J. & Roche, D. B. (2010) Rapid model quality assessment for protein structure predictions using the comparison of multiple models without structural alignments. Bioinformatics, 26, 182-188. PubMed
| Full model quality assessment results for T0585 | Help | ||||
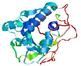
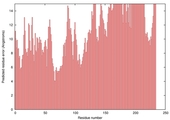
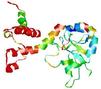
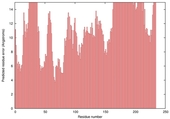
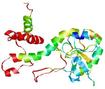
| Model name (PDBsum links for templates used) | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | Model coloured by local quality (click images to view models, local errors and target coverage interactively) |
| nFOLD4_multi _HHsearch_TS1 1jwqA 1xovA |
MEDIUM: 1.208E-2 |
0.2510 |
 |
 |
| nFOLD4_1jwqA _COMA_TS1 |
MEDIUM: 1.395E-2 |
0.2429 |
 |
 |
| nFOLD4_1jwqa _spk2_TS1 |
MEDIUM: 1.493E-2 |
0.2391 |
 |
 |
| nFOLD4_1jwqa _sp3_TS1 |
MEDIUM: 1.493E-2 |
0.2391 |
 |
 |
| nFOLD4_1xovA _HHsearch_TS2 |
MEDIUM: 1.637E-2 |
0.2339 |
 |
 |
| nFOLD4_1xovA _COMA_TS3 |
MEDIUM: 2.06E-2 |
0.2215 |
 |
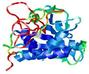 |
| nFOLD4_3czxA _HHsearch_TS3 |
MEDIUM: 2.113E-2 |
0.2202 |
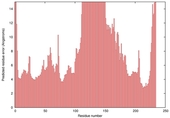 |
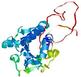 |
| nFOLD4_3czxA _COMA_TS2 |
MEDIUM: 2.161E-2 |
0.2190 |
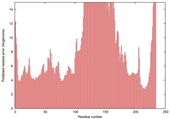 |
 |
| nFOLD4_1jwqA _HHsearch_TS1 |
MEDIUM: 2.228E-2 |
0.2174 |
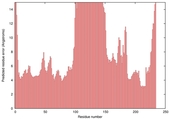 |
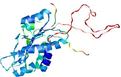 |
| nFOLD4_1xova _sp3_TS2 |
MEDIUM: 2.45E-2 |
0.2124 |
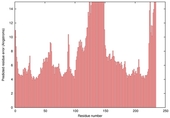 |
 |
| nFOLD4_3czxa _spk2_TS3 |
MEDIUM: 2.653E-2 |
0.2084 |
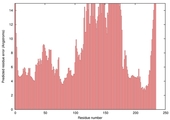 |
 |
| nFOLD4_1xova _spk2_TS2 |
MEDIUM: 2.805E-2 |
0.2055 |
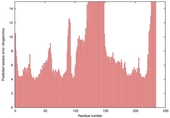 |
 |
| nFOLD4_3czxa _sp3_TS3 |
MEDIUM: 3.23E-2 |
0.1984 |
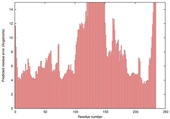 |
 |
| nFOLD4_1xu7a _sp3_TS4 |
POOR: 1.523E-1 |
0.1282 |
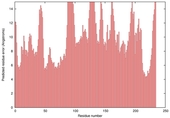 |
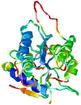 |
| nFOLD4_1yb1a _sp3_TS8 |
POOR: 1.541E-1 |
0.1276 |
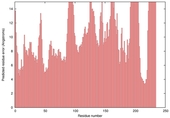 |
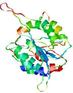 |
| nFOLD4_1euaa _sp3_TS10 |
POOR: 1.772E-1 |
0.1218 |
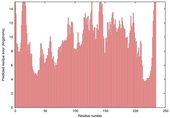 |
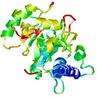 |
| nFOLD4_1euaa _spk2_TS6 |
POOR: 1.958E-1 |
0.1177 |
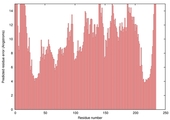 |
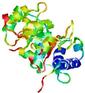 |
| nFOLD4_2v81a _spk2_TS10 |
POOR: 2.127E-1 |
0.1143 |
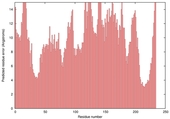 |
 |
| nFOLD4_2b4qa _sp3_TS6 |
POOR: 2.386E-1 |
0.1097 |
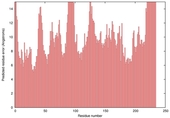 |
 |
| nFOLD4_3cxra _sp3_TS9 |
POOR: 2.398E-1 |
0.1094 |
 |
 |
| nFOLD4_3e9na _spk2_TS9 |
POOR: 3.033E-1 |
0.0998 |
 |
 |
| nFOLD4_2ek8a _sp3_TS5 |
POOR: 3.259E-1 |
0.0969 |
 |
 |
| nFOLD4_2odfa _spk2_TS7 |
POOR: 3.376E-1 |
0.0954 |
 |
 |
| nFOLD4_1wdna _spk2_TS5 |
POOR: 3.642E-1 |
0.0923 |
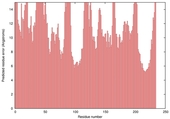 |
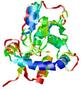 |
| nFOLD4_1lst_ _spk2_TS4 |
POOR: 3.651E-1 |
0.0922 |
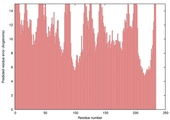 |
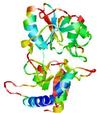 |
| nFOLD4_3dfua _spk2_TS8 |
POOR: 4.157E-1 |
0.0868 |
 |
 |
| nFOLD4_3dfua _sp3_TS7 |
POOR: 4.246E-1 |
0.0859 |
 |
 |
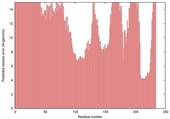
| nFOLD4_1wwpA _HHsearch_TS7 |
POOR: 6.544E-1 |
0.0660 |
 |
 |
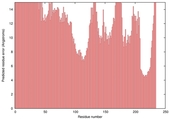
| nFOLD4_1wtyA _HHsearch_TS8 |
POOR: 6.563E-1 |
0.0659 |
 |
 |
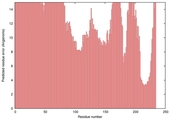
| nFOLD4_1jogA _HHsearch_TS6 |
POOR: 6.684E-1 |
0.0649 |
 |
 |
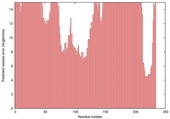
| nFOLD4_2b3wA _HHsearch_TS5 |
POOR: 7.156E-1 |
0.0612 |
 |
 |
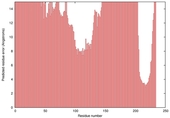
| nFOLD4_3cwvA _HHsearch_TS9 |
POOR: 7.829E-1 |
0.0558 |
 |
 |
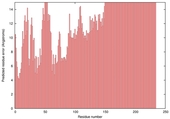
| nFOLD4_1dquA _HHsearch_TS10 |
POOR: 8.546E-1 |
0.0496 |
 |
 |
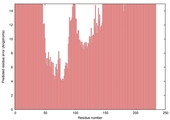
| nFOLD4_3menA _HHsearch_TS4 |
POOR: 9.173E-1 |
0.0429 |
 |
 |