MultiFOLD: automated prediction, quality assessment and refinement of tertiary and quaternary structure models of proteins
About the server
The MultiFOLD server provides:
- State-of-the-art prediction of both tertiary and quaternary structures of proteins.
- Integrated predictions of both global and local model quality for the top refined models.
- Interactive 3D views of predicted multimeric assemblies which can be coloured by chain ID or accuracy.
- Machine-readable data downloads.
This website is free and open to all users and there is no login requirement.
Privacy and cookies
Fair usage policy:
You are only permitted to have 1 job running at a time for each IP address, so please wait until your previous job completes before submitting further data. If you already have a job running then you will be notified and your uploaded data will be deleted. Once your job has completed your IP address will be unlocked and you will be able to submit new data.
MultiFOLD server submission form
Help page
Sample output
References
The MultiFOLD server is described in this paper:- McGuffin, L. J., Edmunds N. S., Genc, A. G., Alharbi, S. M. A., Salehe, B. R. and Adiyaman, R. (2023) Prediction of protein structures, functions and interactions using the IntFOLD7, MultiFOLD and ModFOLDdock servers. Nucleic Acids Research. gkad297. DOI PubMed
- Adiyaman, R., Edmunds, N. S., Genc, A. G., Alharbi, S. M. A. and McGuffin, L. J. (2023) Improvement of protein tertiary and quaternary structure predictions using the ReFOLD refinement method and the AlphaFold2 recycling process. Bioinformatics Advances, vbad078. DOI
Docker package
The Docker package for MultiFOLD, MultiFOLD_refine and ModFOLDdock is available here: https://hub.docker.com/r/mcguffin/multifold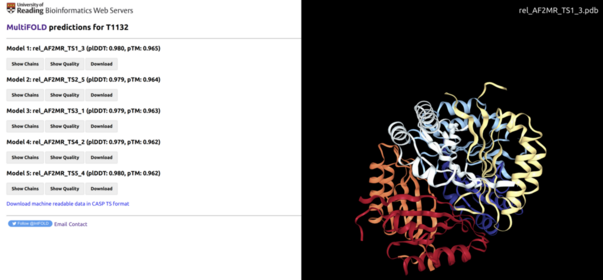
Example of the desktop view of the MultiFOLD results page for CASP target T1132
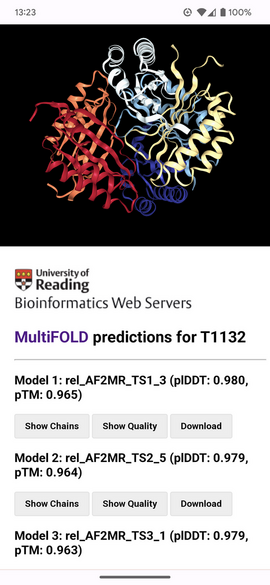
Example of the mobile (portrait) view of the MultiFOLD results page for CASP target T1132