The ModFOLD8 Server Results
Please cite the ModFOLD8 paper:McGuffin, L.J., Aldowsari, F., Alharbi, S. and Adiyaman, R. (2021) ModFOLD8: accurate global and local quality estimates for 3D protein models. Nucleic Acids Research, 49, W425-W430. DOI PubMed
Click on the links below to download machine readable results files:
- Download Quality Assessment file (raw model quality prediction scores in CASP QA format)
- Make a compressed file containing all evaluated 3D models (models are in PDB format with predicted per-residue errors in the B-factor column)
| Graphical ModFOLD8 results for T1024 | Help | |||||
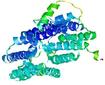
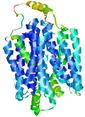
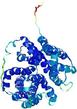
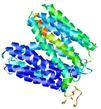
| Model rank | Model ID | Confidence and P-value | Global model quality score | Residue error plot | 3D view of residue error |
| 1 | server15_TS1 |
CERT: 7.103E-16 |
0.6694 | 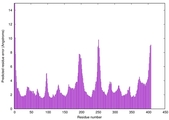
|

|
| 2 | server11_TS1 |
CERT: 4.281E-13 |
0.6412 | 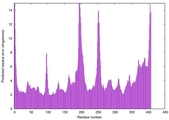
|
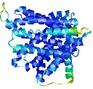
|
| 3 | server14_TS1 |
CERT: 1.763E-11 |
0.6248 | 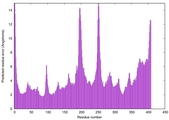
|
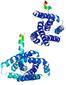
|
| 4 | server13_TS1 |
CERT: 5.398E-7 |
0.5793 | 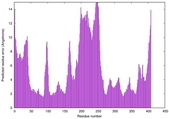
|
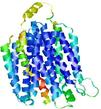
|
| 5 | server16_TS1 |
CERT: 2.88E-6 |
0.5720 | 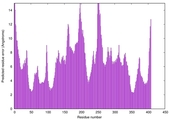
|
|
| 6 | server19_TS1 |
CERT: 6.272E-4 |
0.5482 | 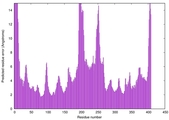
|

|
| 7 | server12_TS1 |
CERT: 6.592E-4 |
0.5480 | 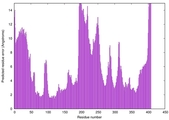
|
|
| 8 | server20_TS1 |
HIGH: 2.898E-3 |
0.5376 | 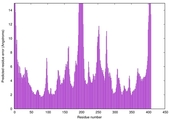
|

|
| 9 | server18_TS1 |
HIGH: 3.489E-3 |
0.5360 | 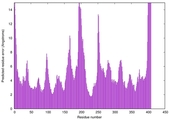
|
|
| 10 | server01_TS1 |
HIGH: 5.215E-3 |
0.5322 | 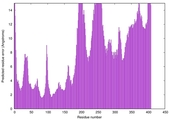
|

|
| 11 | server17_TS1 |
MEDIUM: 1.001E-2 |
0.5242 | 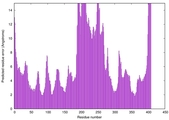
|
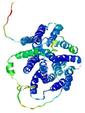
|
| 12 | server05_TS1 |
MEDIUM: 2.652E-2 |
0.5067 | 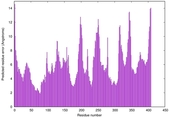
|
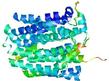
|
| 13 | server10_TS1 |
MEDIUM: 4.413E-2 |
0.4937 | 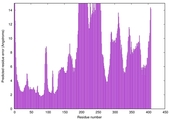
|
|
| 14 | server09_TS1 |
MEDIUM: 4.961E-2 |
0.4902 | 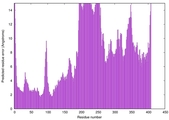
|

|
| 15 | server02_TS1 |
LOW: 5.071E-2 |
0.4895 | 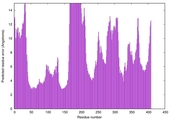
|
|
| 16 | server07_TS1 |
LOW: 5.508E-2 |
0.4869 | 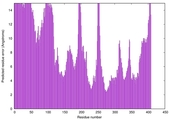
|
|
| 17 | server04_TS1 |
LOW: 5.901E-2 |
0.4853 | 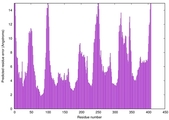
|
|
| 18 | server08_TS1 |
UNCERT: 1.26E-1 |
0.4579 | 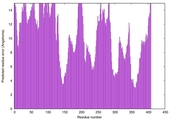
|
|
| 19 | server03_TS1 |
UNCERT: 1.268E-1 |
0.4576 | 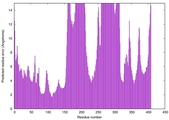
|
|
| 20 | server06_TS1 |
UNCERT: 2.948E-1 |
0.3989 | 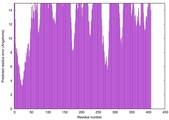
|

|