The IntFOLD Server Results Page (Version 7.0)
Please cite the relevant paper:McGuffin, L. J., Edmunds N. S., Genc, A. G., Alharbi, S. M. A., Salehe, B. R. and Adiyaman, R. (2023) Prediction of protein structures, functions and interactions using the IntFOLD7, MultiFOLD and ModFOLDdock servers. Nucleic Acids Research. gkad297. DOI PubMed
Links to graphical output:
- Top 5 3D models
- Disorder prediction
- Domain boundary prediction
- Binding site prediction
- Full model quality assessment results
Download machine readable results in CASP format:
- TS (Tertiary Structure Prediction)
- DR (Disorder Prediction)
- DP (Domain Prediction)
- FN (Binding Site Prediction)
- QA (Model Quality Prediction)
- Make a compressed file containing all generated 3D models (models are in PDB format with predicted per-residue errors in the B-factor column)
Results will be available for 21 days after job completion (subject to server capacity)
| Tertiary Structure Predictions: Top 5 multi-template 3D models for T1124 | Help | ||||
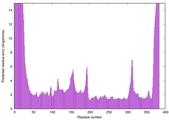
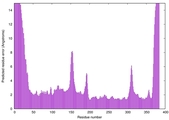
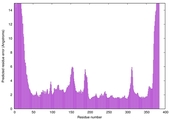
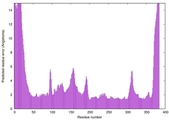
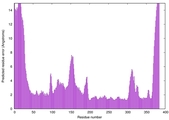
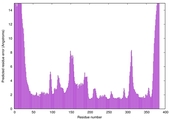
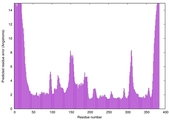
| Model ID and PDBsum links for all templates used | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | 3D map of model quality |
|
IntFOLD7 AF2 TS2 2 0 rel |
CERT: 3.115E-5 |
0.7433 |
 |

|
|
IntFOLD7 AF2 TS4 1 0 rel |
CERT: 3.437E-5 |
0.7381 |
 |

|
|
IntFOLD7 AF2 TS1 3 0 rel |
CERT: 3.617E-5 |
0.7353 |
 |

|
|
IntFOLD7 AF2 TS5 4 0 rel |
CERT: 3.737E-5 |
0.7336 |
 |

|
|
IntFOLD7 AF2 TS4 1 0 |
CERT: 3.929E-5 |
0.7309 |
 |

|
McGuffin, L. J. (2008) Intrinsic disorder prediction from the analysis of multiple protein fold recognition models. Bioinformatics, 24, 1789-1804. PubMed
| Disorder prediction for T1124 | Help | |||
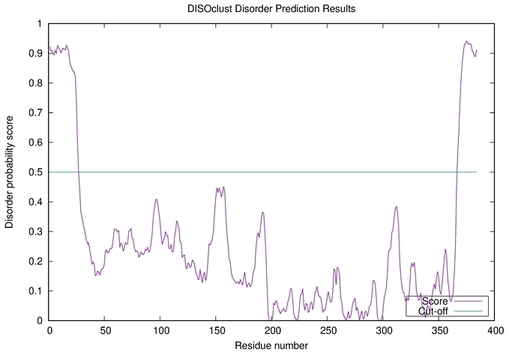
| |||
| Click image to download plot in PostScript format. |
| Domain boundary prediction for T1124 | Help | |||

| |||
| Click image to download model or view prediction using Jmol. The model above is coloured according to the predicted domains. |
Roche, D. B., Buenavista, M. T., McGuffin, L. J. (2013) The FunFOLD2 server for the prediction of protein-ligand interactions. Nucleic Acids Res., 41, W303-7. PubMed
| Ligand binding residues prediction for T1124 | Help | |||

| |||
| Click image to download model or view prediction using Jmol. Predicted ligand binding residues are shown as blue sticks in the image above. |
McGuffin, L.J., Aldowsari, F., Alharbi, S. and Adiyaman, R. (2021) ModFOLD8: accurate global and local quality estimates for 3D protein models. Nucleic Acids Research, 49, W425-W430. DOI PubMed
| Full model quality assessment results for T1124 | Help | ||||
| Model ID and PDBsum links for all templates used | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | 3D map of model quality |
|
IntFOLD7 AF2 TS2 2 0 rel |
CERT: 3.115E-5 |
0.7433 |
 |

|
|
IntFOLD7 AF2 TS4 1 0 rel |
CERT: 3.437E-5 |
0.7381 |
 |

|
|
IntFOLD7 AF2 TS1 3 0 rel |
CERT: 3.617E-5 |
0.7353 |
 |

|
|
IntFOLD7 AF2 TS5 4 0 rel |
CERT: 3.737E-5 |
0.7336 |
 |

|
|
IntFOLD7 AF2 TS4 1 0 |
CERT: 3.929E-5 |
0.7309 |
 |

|
|
IntFOLD7 AF2 TS3 5 0 rel |
CERT: 3.946E-5 |
0.7307 |
 |

|
|
IntFOLD7 AF2 TS2 2 0 |
CERT: 4.204E-5 |
0.7273 |
 |

|
|
IntFOLD7 AF2 TS3 5 0 |
CERT: 5.123E-5 |
0.7168 |
 |

|
|
IntFOLD7 AF2 TS5 4 0 |
CERT: 5.632E-5 |
0.7118 |
 |

|
|
IntFOLD7 AF2 TS1 3 0 |
CERT: 6.279E-5 |
0.7060 |
 |

|
|
IntFOLD7 tR2 TS0 1 0.35 |
CERT: 1.073E-4 |
0.6775 |
 |

|
|
IntFOLD7 tR2 TS0 1 0.25 |
CERT: 1.386E-4 |
0.6638 |
 |

|
|
IntFOLD7 tR2 TS0 0 0.35 |
CERT: 1.469E-4 |
0.6607 |
 |

|
|
IntFOLD7 tR2 TS0 0 0.05 |
CERT: 1.547E-4 |
0.6580 |
 |

|
|
IntFOLD7 tR2 TS0 2 0.35 |
CERT: 1.595E-4 |
0.6564 |
 |

|
|
IntFOLD7 tR2 TS0 1 0.15 |
CERT: 1.692E-4 |
0.6532 |
 |

|
|
IntFOLD7 tR2 TS0 1 0.05 |
CERT: 1.755E-4 |
0.6513 |
 |

|
|
IntFOLD7 tR2 TS0 2 0.05 |
CERT: 1.914E-4 |
0.6467 |
 |

|
|
IntFOLD7 tR2 TS0 0 0.25 |
CERT: 2.087E-4 |
0.6421 |
 |

|
|
IntFOLD7 tR2 TS0 0 0.45 |
CERT: 2.285E-4 |
0.6372 |
 |

|
|
IntFOLD7 tR2 TS0 2 0.15 |
CERT: 2.438E-4 |
0.6338 |
 |

|
|
IntFOLD7 tR2 TS0 0 0.15 |
CERT: 2.799E-4 |
0.6264 |
 |

|
|
IntFOLD7 tR2 TS0 1 0.45 |
CERT: 2.811E-4 |
0.6262 |
 |

|
|
IntFOLD7 tR2 TS0 2 0.25 |
CERT: 2.909E-4 |
0.6244 |
 |

|
|
IntFOLD7 tR2 TS0 2 0.45 |
CERT: 3.122E-4 |
0.6206 |
 |

|
|
IntFOLD7 dPPAS multi3 TS1 3mczA 4qvgA 2r3sA 2ip2A 3gwzA 4a6dA |
CERT: 5.38E-4 |
0.5917 |
 |

|
|
IntFOLD7 HHpred TS2 3mczA 2r3sA 5i2hA 4a6dA 4z2yA |
CERT: 5.82E-4 |
0.5875 |
 |

|
|
IntFOLD7 HHpred TS1 3mczA 2r3sA 5i2hA 4a6dA 4z2yA |
CERT: 5.902E-4 |
0.5867 |
 |

|
|
IntFOLD7 HHpred TS3 3mczA 2r3sA 5i2hA 4a6dA 4z2yA |
CERT: 6.101E-4 |
0.5850 |
 |

|
|
IntFOLD7 IntFOLDTS60 multi4 TS1 3gwzA 4a6dA 3mczA 2r3sA 1x19A |
CERT: 6.354E-4 |
0.5828 |
 |

|
|
IntFOLD7 dPPAS2 multi7 TS1 3gwzA 4a6dA |
CERT: 6.664E-4 |
0.5803 |
 |

|
|
IntFOLD7 dPPAS2 multi5 TS1 3gwzA 4a6dA |
CERT: 6.664E-4 |
0.5803 |
 |

|
|
IntFOLD7 wMUSTER multi8 TS1 7clfA 4a6dA 1tw2B 3mczA 4qvgA |
CERT: 6.674E-4 |
0.5802 |
 |

|
|
IntFOLD7 IntFOLDTS140 multi4 TS1 3gwzA 2r3sA 4a6dA 3mczA 4qvgA |
CERT: 6.693E-4 |
0.5800 |
 |

|
|
IntFOLD7 SPARKSX multi8 TS1 7clfA 2r3sA 3gwzD 3mczA 1x19A 5i2hA |
CERT: 6.746E-4 |
0.5796 |
 |

|
|
IntFOLD7 CNFsearch multi5 TS1 4a6dA 3gwzA |
CERT: 6.804E-4 |
0.5792 |
 |

|
|
IntFOLD7 LOMETS multi3 TS1 3mczA 7clfA |
CERT: 6.838E-4 |
0.5789 |
 |

|
|
IntFOLD7 Env-PPAS multi3 TS1 3mczA 7clfA |
CERT: 7.038E-4 |
0.5774 |
 |

|
|
IntFOLD7 Env-PPAS multi1 TS1 3mczA 7clfA |
CERT: 7.038E-4 |
0.5774 |
 |

|
|
IntFOLD7 IntFOLDTS60 multi3 TS1 3gwzA 4a6dA |
CERT: 7.178E-4 |
0.5763 |
 |

|
|
IntFOLD7 IntFOLDTS60 multi1 TS1 3gwzA 4a6dA |
CERT: 7.178E-4 |
0.5763 |
 |
|
|
IntFOLD7 IntFOLDTS140 multi3 TS1 3gwzA 4a6dA |
CERT: 7.178E-4 |
0.5763 |
 |

|
|
IntFOLD7 HHsearch multi7 TS1 3gwzA 4a6dA |
CERT: 7.178E-4 |
0.5763 |
 |

|
|
IntFOLD7 HHsearch multi5 TS1 3gwzA 4a6dA |
CERT: 7.178E-4 |
0.5763 |
 |

|
|
IntFOLD7 sp3 multi8 TS1 3gwzA 4a6dA 3mczA 1x19A |
CERT: 8.192E-4 |
0.5693 |
 |

|
|
IntFOLD7 sp3 multi7 TS1 3gwzA 4a6dA |
CERT: 8.354E-4 |
0.5682 |
 |

|
|
IntFOLD7 sp3 multi5 TS1 3gwzA 4a6dA |
CERT: 8.354E-4 |
0.5682 |
 |

|
|
IntFOLD7 HHsearch multi8 TS1 3gwzA 4a6dA 7clfA 3mczA |
CERT: 8.442E-4 |
0.5677 |
 |

|
|
IntFOLD7 HHsearch multi3 TS1 4qvgA 7clfA 4a6dA |
CERT: 8.461E-4 |
0.5676 |
 |

|
|
IntFOLD7 PPAS multi3 TS1 3mczA 7clfA |
CERT: 8.577E-4 |
0.5668 |
 |

|
|
IntFOLD7 PPAS multi1 TS1 3mczA 7clfA |
CERT: 8.577E-4 |
0.5668 |
 |

|
|
IntFOLD7 wPPAS multi8 TS1 7clfA 3gwzA 1tw2B 2r3sA 3mczA 1x19A |
CERT: 8.76E-4 |
0.5657 |
 |

|
|
IntFOLD7 wdPPAS multi8 TS1 3gwzA 4a6dA 3mczA 1x19A |
CERT: 9.202E-4 |
0.5631 |
 |

|
|
IntFOLD7 HHsearch multi1 TS1 4qvgA 5i2hA |
CERT: 9.408E-4 |
0.5619 |
 |

|
|
IntFOLD7 COMA multi8 TS1 7clfA 6clwA 5i2hA 4m6xA |
CERT: 9.419E-4 |
0.5619 |
 |

|
|
IntFOLD7 dPPAS2 multi8 TS1 3gwzA 4a6dA 1tw2B 3mczA 1x19A |
CERT: 9.421E-4 |
0.5618 |
 |

|
|
IntFOLD7 wPPAS multi3 TS1 3mczA 7clfA |
CERT: 9.489E-4 |
0.5615 |
 |

|
|
IntFOLD7 wPPAS multi1 TS1 3mczA 7clfA |
CERT: 9.489E-4 |
0.5615 |
 |

|
|
IntFOLD7 sp3 multi3 TS1 3mczA 4a6dA |
CERT: 9.521E-4 |
0.5613 |
 |

|
|
IntFOLD7 sp3 multi1 TS1 3mczA 4a6dA |
CERT: 9.521E-4 |
0.5613 |
 |

|
|
IntFOLD7 dPPAS multi7 TS1 4a6dA |
CERT: 9.828E-4 |
0.5596 |
 |

|
|
IntFOLD7 dPPAS multi6 TS1 4a6dA |
CERT: 9.828E-4 |
0.5596 |
 |

|
|
IntFOLD7 CNFsearch multi1 TS1 2r3sA 5i2hA |
CERT: 9.877E-4 |
0.5593 |
 |

|
|
IntFOLD7 CNFsearch multi3 TS1 2r3sA 3gwzA 6c5bA 4a6dA |
CERT: 9.896E-4 |
0.5592 |
 |

|
|
IntFOLD7 Env-PPAS multi4 TS1 3mczA 5i2hA 1x19A |
CERT: 9.942E-4 |
0.5590 |
 |

|
|
IntFOLD7 IT5 TS2 |
CERT: 9.996E-4 |
0.5587 |
 |

|
|
IntFOLD7 dPPAS2 multi3 TS1 3mczA 2r3sA 3gwzA |
HIGH: 1.002E-3 |
0.5586 |
 |

|
|
IntFOLD7 wMUSTER multi3 TS1 3mczA 7clfA |
HIGH: 1.016E-3 |
0.5578 |
 |

|
|
IntFOLD7 wMUSTER multi1 TS1 3mczA 7clfA |
HIGH: 1.016E-3 |
0.5578 |
 |

|
|
IntFOLD7 SPARKSX multi2 TS1 5i2hA |
HIGH: 1.03E-3 |
0.5571 |
 |

|
|
IntFOLD7 wPPAS multi7 TS1 7clfA |
HIGH: 1.058E-3 |
0.5557 |
 |

|
|
IntFOLD7 wPPAS multi6 TS1 7clfA |
HIGH: 1.058E-3 |
0.5557 |
 |

|
|
IntFOLD7 SPARKSX multi7 TS1 7clfA |
HIGH: 1.078E-3 |
0.5547 |
 |

|
|
IntFOLD7 SPARKSX multi6 TS1 7clfA |
HIGH: 1.078E-3 |
0.5547 |
 |

|
|
IntFOLD7 wdPPAS multi1 TS1 7clfA 3mczA |
HIGH: 1.093E-3 |
0.5539 |
 |

|
|
IntFOLD7 CNFsearch multi4 TS1 2r3sA 5i2hA 3mczA |
HIGH: 1.099E-3 |
0.5536 |
 |

|
|
IntFOLD7 wMUSTER multi5 TS1 7clfA 4a6dA |
HIGH: 1.101E-3 |
0.5536 |
 |

|
|
IntFOLD7 wMUSTER multi7 TS1 7clfA |
HIGH: 1.106E-3 |
0.5533 |
 |

|
|
IntFOLD7 wMUSTER multi6 TS1 7clfA |
HIGH: 1.106E-3 |
0.5533 |
 |

|
|
IntFOLD7 IntFOLDTS140 multi1 TS1 3gwzA 2r3sA |
HIGH: 1.113E-3 |
0.5530 |
 |

|
|
IntFOLD7 SPARKSX multi5 TS1 7clfA 2r3sA |
HIGH: 1.115E-3 |
0.5529 |
 |

|
|
IntFOLD7 Env-PPAS multi7 TS1 3gwzA 7clfA |
HIGH: 1.13E-3 |
0.5522 |
 |

|
|
IntFOLD7 Env-PPAS multi5 TS1 3gwzA 7clfA |
HIGH: 1.13E-3 |
0.5522 |
 |

|
|
IntFOLD7 spk2 multi8 TS1 3gwzA 3mczA 1x19A |
HIGH: 1.158E-3 |
0.5509 |
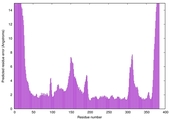 |

|
|
IntFOLD7 PPAS multi7 TS1 3gwzA 7clfA |
HIGH: 1.165E-3 |
0.5506 |
 |

|
|
IntFOLD7 PPAS multi5 TS1 3gwzA 7clfA |
HIGH: 1.165E-3 |
0.5506 |
 |

|
|
IntFOLD7 SPARKSX multi4 TS1 5i2hA 4qvgA 3mczA |
HIGH: 1.183E-3 |
0.5497 |
 |

|
|
IntFOLD7 HHsearch multi4 TS1 4qvgA 5i2hA 3mczA |
HIGH: 1.193E-3 |
0.5493 |
 |

|
|
IntFOLD7 dPPAS multi5 TS1 4a6dA 6c5bA |
HIGH: 1.2E-3 |
0.5489 |
 |

|
|
IntFOLD7 wdPPAS multi3 TS1 7clfA |
HIGH: 1.22E-3 |
0.5481 |
 |

|
|
IntFOLD7 wdPPAS multi2 TS1 7clfA |
HIGH: 1.22E-3 |
0.5481 |
 |

|
|
IntFOLD7 dPPAS multi1 TS1 3mczA 4qvgA |
HIGH: 1.221E-3 |
0.5481 |
 |

|
|
IntFOLD7 spk2 multi3 TS1 3mczA 3gwzA |
HIGH: 1.261E-3 |
0.5463 |
 |

|
|
IntFOLD7 dPPAS multi8 TS1 4a6dA 1tw2B 1x19A |
HIGH: 1.265E-3 |
0.5462 |
 |

|
|
IntFOLD7 IT5 TS1 |
HIGH: 1.273E-3 |
0.5458 |
 |

|
|
IntFOLD7 sp3 multi6 TS1 3gwzA |
HIGH: 1.275E-3 |
0.5458 |
 |

|
|
IntFOLD7 wdPPAS multi7 TS1 3gwzA 7clfA |
HIGH: 1.278E-3 |
0.5456 |
 |

|
|
IntFOLD7 wdPPAS multi5 TS1 3gwzA 7clfA |
HIGH: 1.278E-3 |
0.5456 |
 |

|
|
IntFOLD7 spk2 multi4 TS1 3mczA 2r3sA 1x19A |
HIGH: 1.285E-3 |
0.5453 |
 |

|
|
IntFOLD7 CNFsearch multi2 TS1 2r3sA |
HIGH: 1.285E-3 |
0.5453 |
 |

|
|
IntFOLD7 IntFOLDTS60 multi2 TS1 3gwzA |
HIGH: 1.305E-3 |
0.5445 |
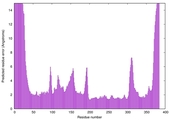 |

|
|
IntFOLD7 IntFOLDTS140 multi2 TS1 3gwzA |
HIGH: 1.305E-3 |
0.5445 |
 |

|
|
IntFOLD7 HHsearch multi6 TS1 3gwzA |
HIGH: 1.305E-3 |
0.5445 |
 |

|
|
IntFOLD7 wPPAS multi4 TS1 3mczA 2r3sA 1x19A |
HIGH: 1.306E-3 |
0.5445 |
 |

|
|
IntFOLD7 COMA multi1 TS1 4qvgA 5i2hA |
HIGH: 1.317E-3 |
0.5440 |
 |

|
|
IntFOLD7 wPPAS multi5 TS1 7clfA 3gwzA |
HIGH: 1.333E-3 |
0.5434 |
 |

|
|
IntFOLD7 PPAS multi8 TS1 3gwzA 1tw2B 3mczA 1x19A |
HIGH: 1.342E-3 |
0.5430 |
 |

|
|
IntFOLD7 CNFsearch multi7 TS1 4a6dA |
HIGH: 1.345E-3 |
0.5429 |
 |

|
|
IntFOLD7 CNFsearch multi6 TS1 4a6dA |
HIGH: 1.345E-3 |
0.5429 |
 |

|
|
IntFOLD7 dPPAS2 multi1 TS1 3mczA 4qvgA |
HIGH: 1.349E-3 |
0.5427 |
 |

|
|
IntFOLD7 wdPPAS multi4 TS1 7clfA 3mczA 1x19A |
HIGH: 1.351E-3 |
0.5427 |
 |

|
|
IntFOLD7 Env-PPAS multi6 TS1 3gwzA |
HIGH: 1.393E-3 |
0.5410 |
 |

|
|
IntFOLD7 wMUSTER multi4 TS1 3mczA 4qvgA |
HIGH: 1.481E-3 |
0.5378 |
 |

|
|
IntFOLD7 wdPPAS multi6 TS1 3gwzA |
HIGH: 1.504E-3 |
0.5370 |
 |

|
|
IntFOLD7 Env-PPAS multi8 TS1 3gwzA 1tw2B 3mczA 1x19A |
HIGH: 1.541E-3 |
0.5357 |
 |

|
|
IntFOLD7 dPPAS2 multi6 TS1 3gwzA |
HIGH: 1.627E-3 |
0.5328 |
 |

|
|
IntFOLD7 spk2 multi5 TS1 3gwzA 3mczA |
HIGH: 1.691E-3 |
0.5307 |
 |

|
|
IntFOLD7 COMA multi3 TS1 4qvgA 7clfA |
HIGH: 1.741E-3 |
0.5292 |
 |

|
|
IntFOLD7 IT5 TS3 |
HIGH: 1.758E-3 |
0.5287 |
 |

|
|
IntFOLD7 COMA multi4 TS1 4qvgA 5i2hA 4m6xA 4d7kA |
HIGH: 1.78E-3 |
0.5280 |
 |

|
|
IntFOLD7 COMA multi7 TS1 7clfA |
HIGH: 1.821E-3 |
0.5268 |
 |

|
|
IntFOLD7 COMA multi6 TS1 7clfA |
HIGH: 1.821E-3 |
0.5268 |
 |

|
|
IntFOLD7 PPAS multi4 TS1 3mczA 1x19A |
HIGH: 1.883E-3 |
0.5250 |
 |

|
|
IntFOLD7 COMA multi5 TS1 7clfA 6clwA |
HIGH: 1.9E-3 |
0.5245 |
 |

|
|
IntFOLD7 PPAS multi6 TS1 3gwzA |
HIGH: 1.922E-3 |
0.5239 |
 |

|
|
IntFOLD7 CNFsearch multi8 TS1 4a6dA 2ip2A |
HIGH: 1.922E-3 |
0.5239 |
 |

|
|
IntFOLD7 spk2 multi7 TS1 3gwzA |
HIGH: 1.925E-3 |
0.5238 |
 |

|
|
IntFOLD7 spk2 multi6 TS1 3gwzA |
HIGH: 1.925E-3 |
0.5238 |
 |

|
|
IntFOLD7 dPPAS2 multi4 TS1 3mczA 1x19A |
HIGH: 2.037E-3 |
0.5208 |
 |

|
|
IntFOLD7 SPARKSX multi3 TS1 5i2hA 7clfA |
HIGH: 2.038E-3 |
0.5208 |
 |

|
|
IntFOLD7 SPARKSX multi1 TS1 5i2hA 7clfA |
HIGH: 2.038E-3 |
0.5208 |
 |

|
|
IntFOLD7 PPAS multi2 TS1 3mczA |
HIGH: 2.044E-3 |
0.5206 |
 |

|
|
IntFOLD7 Env-PPAS multi2 TS1 3mczA |
HIGH: 2.076E-3 |
0.5198 |
 |

|
|
IntFOLD7 LOMETS multi4 TS1 3mczA |
HIGH: 2.093E-3 |
0.5194 |
 |

|
|
IntFOLD7 sp3 multi4 TS1 3mczA 1x19A |
HIGH: 2.103E-3 |
0.5191 |
 |

|
|
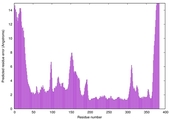
IntFOLD7 spk2 multi1 TS1 3mczA 2r3sA |
HIGH: 2.13E-3 |
0.5184 |
 |

|
|
IntFOLD7 dPPAS multi4 TS1 3mczA 1x19A |
HIGH: 2.133E-3 |
0.5183 |
 |

|
|
IntFOLD7 COMA multi2 TS1 4qvgA |
HIGH: 2.211E-3 |
0.5164 |
 |

|
|
IntFOLD7 HHsearch multi2 TS1 4qvgA |
HIGH: 2.398E-3 |
0.5121 |
 |

|
|
IntFOLD7 sp3 multi2 TS1 3mczA |
HIGH: 2.472E-3 |
0.5105 |
 |

|
|
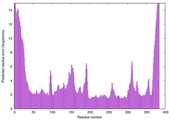
IntFOLD7 wPPAS multi2 TS1 3mczA |
HIGH: 2.54E-3 |
0.5091 |
 |

|
|
IntFOLD7 wMUSTER multi2 TS1 3mczA |
HIGH: 2.939E-3 |
0.5013 |
 |

|
|
IntFOLD7 dPPAS multi2 TS1 3mczA |
HIGH: 3.024E-3 |
0.4998 |
 |

|
|
IntFOLD7 LOMETS multi2 TS1 3mczA |
HIGH: 3.101E-3 |
0.4984 |
 |

|
|
IntFOLD7 dPPAS2 multi2 TS1 3mczA |
HIGH: 3.101E-3 |
0.4984 |
 |

|
|
IntFOLD7 LOMETS multi1 TS1 3mczA |
HIGH: 3.236E-3 |
0.4962 |
 |

|
|
IntFOLD7 spk2 multi2 TS1 3mczA |
HIGH: 3.7E-3 |
0.4890 |
 |

|
|
IntFOLD7 DMPf TS1 |
MEDIUM: 1.035E-2 |
0.4343 |
 |

|
|
IntFOLD7 DMPf TS2 |
MEDIUM: 1.653E-2 |
0.4094 |
 |

|