Links to interactive graphical output:
RasMol generated image of per-residue accuracy for the model nFOLD4_3gwqa_spk2_TS9.bfact.pdb
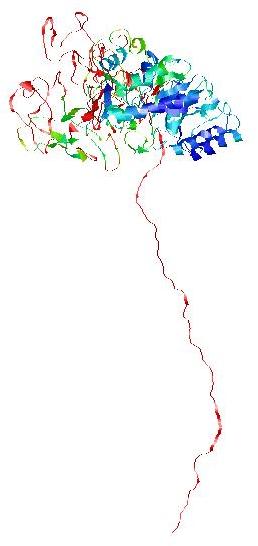
Click here to download a PDB file of this model with residue accuracy predictions (Angstroms) in the B-factor column.
[RasMol colouring uses the reverse rainbow scheme from blue (high accuracy)
through green, yellow and orange to red (low accuracy).]
Jmol view of the per-residue accuracy for the model nFOLD4_3gwqa_spk2_TS9.bfact.pdb
[Jmol colouring uses a gradient from blue (high accuracy)
through white to red (low accuracy).]
Jmol view of the structural alignment of the model (blue) with the template 3gwqA (red) using TM-align
Download the superposition file (RasMol script).
Coverage of target: 0.63829786 (Target length: 611, Aligned residues: 390)
RMSD: 2.11
TM-score: 0.60481
The alignment (template-model) is shown below:
(":" denotes the residue pairs of distance < 5.0 Angstrom)
----ATIDPYSKGLGV--------PGTSIQLTDAARLEWNLLNEDVSLPAAVLYADRVEHNLKWQA-----FVAEYGVK-----LAPHGKTTAPQLFRRQLETGAWG-ITL---ATAHQVRAAYHGG-VSRVLANQLVGRRN-V-AELLSDPEFEFFCLVDSVEGVEQLGEFFK-SVNKQ--LQVLLEL---------GVPGG-RTGVRDAAQRNAVLEAITRYPD-TLKLAGVELYEGV-LKEEHEVREFLQSAVAVTRELVEQERFARAPAVLSGA-------GSAWY--------------D-VVAEEFVKASETGK-VEVVL-RPG-------------CYL------------------------------------TH-D-------VGIYRKAQ-----TDIF-EGLLPALQLWAYVQS-IPEPD-----------RAIIGLGK-----RDSAFDAGPEPARHYRPGNEAPR-------DIAASEGWEIFGLDQHAYLRI--PAGADLKVGDIAFDISHPCLTFDKWR--------QVLVVDP---------AYRVTEVIETFF-----------------------------------------------
:::::::::::: ::.::::::: ::::::::::::::::::::::::::::: :::::::: ::::::::::::::::::::::: ::: ::::::::::::: :::::::::::::: : ::::::::::::::::::::::::::::: ::::: ::::::: ::::: :::::::::::::::::::::: :::: :::::::: :::::::::::::::::::::: :::::::::::::: ::::: : :::::::::::::: ::::: ::: ::: :: : :::::::. :::. ::::::::::::::: ::::. .::::::: :::::: :::::::::::::::: ::::::::::::::::::::: ::::::: :::::::::::::::::. ::::::: ::::::::::::
MMDYGIDIWGNENFIIKNGKVCINYEKKPAIIDI---VKELRDDGYKGPLLLRFPHLIQKQIENIYGNFNKARKEFGYKGGFNAVYPLKVNQYPGFVKNLVKLGKDYNYGLEAGSKAELLLAMAYNNEGAPITVNGFKDRELINIGFIAAEMGHNITLTIEGLNELEAIIDIAKERFKPKPNIGLRVRLHSAGVGIWAKSGGINSKFGLTSTELIEAVNLLKENKLLEQFT-MIHFHLGSQITEIHPLKKALNEAGNIYTELR-KMGAKNLKAINLGGGLAVEYSQFKNEKSRNYTLREYANDVVFILKNIAEQKKDLEPDIFIESGRFVAANHAVLIAPVLELFSQEYAENKLILKKQNPKLIDELYDLYKSIKPSNALEYLHDSIDHLESILTLFDLGYVDLQDRSNAEILTHLITKKAILLLGDKQNPADLLAIQDEVQERYLVNFSLFQSMPDFWGLE-QNFPIMPLDRLDEEPTRSASIWDITCDSDGEISYSKDKPLFLHDVDVEKENYF-LGFFLVGAYQEVLGMKHNLFTHPTEAIISINEKGYEVEGIIEAQSILDTLEDLDYDIHAIMDILNERISNSKLVNDKQKKHILGELYLFLNDNGYLKSIGV
