The IntFOLD Server Results Page (Version 6.0)
Please cite the relevant papers:McGuffin, L.J., Adiyaman, R., Maghrabi, A.H.A., Shuid, A.N., Brackenridge, D.A., Nealon, J.O. & Philomina, L.S. (2019) IntFOLD: an integrated web resource for high performance protein structure and function prediction. Nucleic Acids Research, 47, W408-W413. DOI PubMed
McGuffin, L.J., Shuid, A.M., Kempster, R., Maghrabi, A.H.A., Nealon J.O., Salehe, B.R., Atkins, J.D. & Roche, D.B. (2017) Accurate Template Based Modelling in CASP12 using the IntFOLD4-TS, ModFOLD6 and ReFOLD methods. Proteins: Structure, Function, and Bioinformatics, 86 Suppl 1, 335-344, doi: 10.1002/prot.25360. PubMed
Links to graphical output:
- Top 5 3D models
- Disorder prediction
- Domain boundary prediction
- Binding site prediction
- Full model quality assessment results
Download machine readable results in CASP format:
- TS (Tertiary Structure Prediction)
- DR (Disorder Prediction)
- DP (Domain Prediction)
- FN (Binding Site Prediction)
- QA (Model Quality Prediction)
- Make a compressed file containing all generated 3D models (models are in PDB format with predicted per-residue errors in the B-factor column)
Results will be available for 21 days after job completion (subject to server capacity)
| Tertiary Structure Predictions: Top 5 multi-template 3D models for striatin | Help | ||||
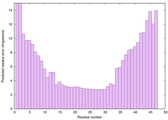
| Model ID and PDBsum links for all templates used | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | 3D map of model quality |
|
IntFOLD6 wdPPAS multi8 TS1 3he5A 5vr2A |
CERT: 1.395E-17 |
0.6867 |
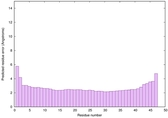 |

|
|
IntFOLD6 wdPPAS multi5 TS1 3he5A 5vr2A |
CERT: 1.395E-17 |
0.6867 |
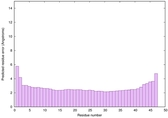 |

|
|
IntFOLD6 wdPPAS multi3 TS1 4n6jA 5vr2A 2oqqA 4pxjA 5al6A |
CERT: 1.71E-17 |
0.6858 |
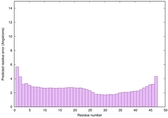 |

|
|
IntFOLD6 wMUSTER multi7 TS1 6h9mA2 3he5A 2oqqA |
CERT: 2.827E-17 |
0.6836 |
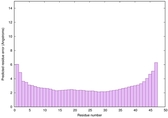 |

|
|
IntFOLD6 MUSTER multi7 TS1 6h9mA2 3he5A 2oqqA |
CERT: 2.827E-17 |
0.6836 |
 |

|
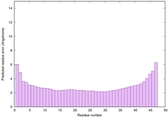
McGuffin, L. J. (2008) Intrinsic disorder prediction from the analysis of multiple protein fold recognition models. Bioinformatics, 24, 1789-1804. PubMed
| Disorder prediction for striatin | Help | |||
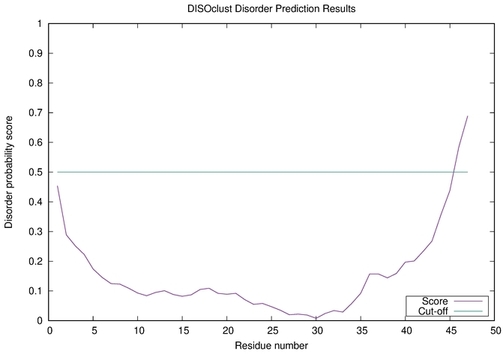
| |||
| Click image to download plot in PostScript format. |
| Domain boundary prediction for striatin | Help | |||

| |||
| Click image to download model or view prediction using Jmol. The model above is coloured according to the predicted domains. |
Roche, D. B., Tetchner, S. J. & McGuffin, L. J. (2011) FunFOLD: an improved automated method for the prediction of ligand binding residues using 3D models of proteins. BMC Bioinformatics, 12, 160. PubMed
| Ligand binding residues prediction for striatin | Help | |||
| |||
| No binding residues predicted. |
Maghrabi, A.H.A. & McGuffin L.J. (2017) ModFOLD6: an accurate web server for the global and local quality estimation of 3D models of proteins. Nucleic Acids Res., 45, W416-W421, doi: 10.1093/nar/gkx332. PubMed
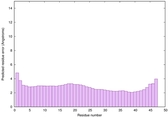
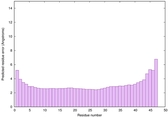
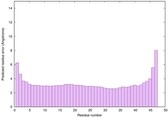
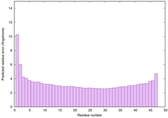
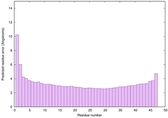
| Full model quality assessment results for striatin | Help | ||||
| Model ID and PDBsum links for all templates used | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | 3D map of model quality |
|
IntFOLD6 wdPPAS multi8 TS1 3he5A 5vr2A |
CERT: 1.395E-17 |
0.6867 |
 |

|
|
IntFOLD6 wdPPAS multi5 TS1 3he5A 5vr2A |
CERT: 1.395E-17 |
0.6867 |
 |

|
|
IntFOLD6 wdPPAS multi3 TS1 4n6jA 5vr2A 2oqqA 4pxjA 5al6A |
CERT: 1.71E-17 |
0.6858 |
 |

|
|
IntFOLD6 wMUSTER multi7 TS1 6h9mA2 3he5A 2oqqA |
CERT: 2.827E-17 |
0.6836 |
 |

|
|
IntFOLD6 MUSTER multi7 TS1 6h9mA2 3he5A 2oqqA |
CERT: 2.827E-17 |
0.6836 |
 |

|
|
IntFOLD6 IT5 TS1 |
CERT: 1.011E-16 |
0.6780 |
 |

|
|
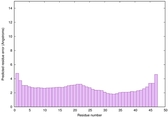
IntFOLD6 CNFsearch multi4 TS1 4n6jA 4wheA |
CERT: 1.148E-16 |
0.6774 |
 |

|
|
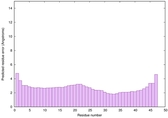
IntFOLD6 CNFsearch multi3 TS1 4n6jA 4wheA |
CERT: 1.148E-16 |
0.6774 |
 |

|
|
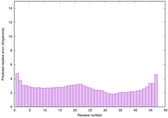
IntFOLD6 CNFsearch multi1 TS1 4n6jA 4wheA |
CERT: 1.148E-16 |
0.6774 |
 |

|
|
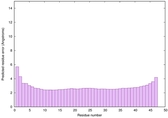
IntFOLD6 dPPAS2 multi8 TS1 4pkyB 6h9mA2 |
CERT: 1.397E-16 |
0.6766 |
 |

|
|
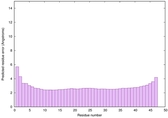
IntFOLD6 dPPAS2 multi5 TS1 4pkyB 6h9mA2 |
CERT: 1.397E-16 |
0.6766 |
 |

|
|
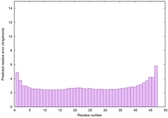
IntFOLD6 IntFOLDTS60 multi4 TS1 4n6jA 6f1tX |
CERT: 2.244E-16 |
0.6745 |
 |

|
|
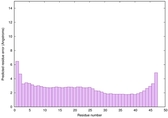
IntFOLD6 IntFOLDTS140 multi2 TS1 2x7aA |
CERT: 2.279E-16 |
0.6744 |
 |

|
|
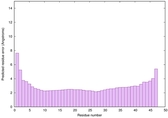
IntFOLD6 dPPAS multi3 TS1 4n6jA 5vr2A 2oqqA 3he5A 5al6A 5kb0A |
CERT: 2.388E-16 |
0.6742 |
 |

|
|
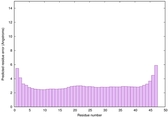
IntFOLD6 HHpred TS1 4n6jA |
CERT: 4.117E-16 |
0.6718 |
 |

|
|
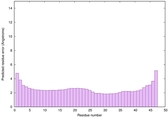
IntFOLD6 IntFOLDTS60 multi3 TS1 4n6jA 4ywzA 4zqaA 2p4vA |
CERT: 5.823E-16 |
0.6703 |
 |

|
|
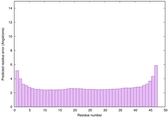
IntFOLD6 IntFOLDTS60 multi1 TS1 4n6jA |
CERT: 5.985E-16 |
0.6702 |
 |

|
|
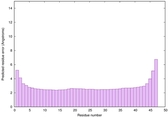
IntFOLD6 HHpred TS2 4n6jA |
CERT: 7.32E-16 |
0.6693 |
 |

|
|
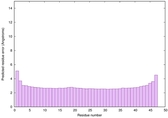
IntFOLD6 dPPAS2 multi7 TS1 4pkyB 3he5A 4pxjA |
CERT: 9.825E-16 |
0.6680 |
 |

|
|
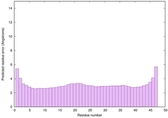
IntFOLD6 HHsearch multi6 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
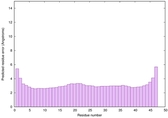
IntFOLD6 HHsearch multi2 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
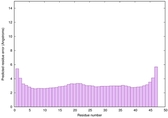
IntFOLD6 COMA multi8 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
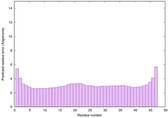
IntFOLD6 COMA multi7 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
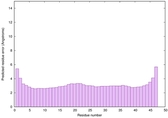
IntFOLD6 COMA multi6 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
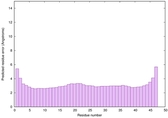
IntFOLD6 COMA multi5 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
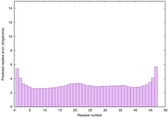
IntFOLD6 COMA multi4 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
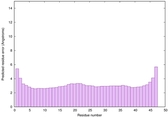
IntFOLD6 COMA multi3 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
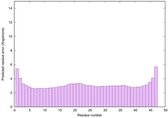
IntFOLD6 COMA multi2 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
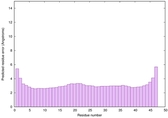
IntFOLD6 COMA multi1 TS1 4n6jA |
CERT: 1.036E-15 |
0.6677 |
 |

|
|
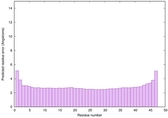
IntFOLD6 wPPAS multi3 TS1 4n6jA 5vr2A 2oqqA 4pxjA |
CERT: 1.246E-15 |
0.6669 |
 |

|
|
IntFOLD6 CNFsearch multi5 TS1 4wheA 4n6jA |
CERT: 1.843E-15 |
0.6652 |
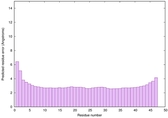 |

|
|
IntFOLD6 SPARKSX multi7 TS1 4n6jA |
CERT: 1.924E-15 |
0.6650 |
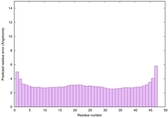 |

|
|
IntFOLD6 SPARKSX multi6 TS1 4n6jA |
CERT: 1.924E-15 |
0.6650 |
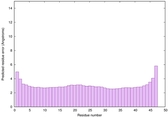 |

|
|
IntFOLD6 SPARKSX multi3 TS1 4n6jA |
CERT: 1.924E-15 |
0.6650 |
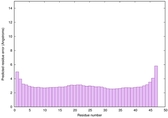 |

|
|
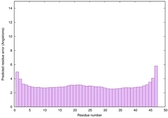
IntFOLD6 SPARKSX multi2 TS1 4n6jA |
CERT: 1.924E-15 |
0.6650 |
 |

|
|
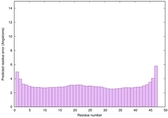
IntFOLD6 IntFOLDTS60 multi2 TS1 4n6jA |
CERT: 1.924E-15 |
0.6650 |
 |

|
|
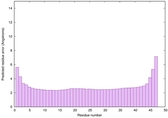
IntFOLD6 HHpred TS3 4n6jA |
CERT: 1.966E-15 |
0.6649 |
 |

|
|
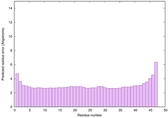
IntFOLD6 HHsearch multi3 TS1 4n6jA 4zqaA 2p4vA |
CERT: 2.695E-15 |
0.6635 |
 |

|
|
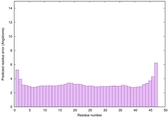
IntFOLD6 CNFsearch multi2 TS1 4n6jA |
CERT: 2.901E-15 |
0.6632 |
 |

|
|
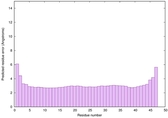
IntFOLD6 IntFOLDTS140 multi4 TS1 2x7aA 6h9mA2 |
CERT: 2.996E-15 |
0.6631 |
 |

|
|
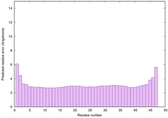
IntFOLD6 IntFOLDTS140 multi1 TS1 2x7aA 6h9mA2 |
CERT: 2.996E-15 |
0.6631 |
 |

|
|
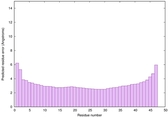
IntFOLD6 dPPAS multi7 TS1 6h9mA2 3he5A 2oqqA 5al6A 5kb0A |
CERT: 3.006E-15 |
0.6630 |
 |

|
|
IntFOLD6 wMUSTER multi5 TS1 6h9mA2 3he5A |
CERT: 3.086E-15 |
0.6629 |
 |

|
|
IntFOLD6 PPAS multi5 TS1 6h9mA2 3he5A |
CERT: 3.086E-15 |
0.6629 |
 |

|
|
IntFOLD6 MUSTER multi5 TS1 6h9mA2 3he5A |
CERT: 3.086E-15 |
0.6629 |
 |

|
|
IntFOLD6 dPPAS multi5 TS1 6h9mA2 3he5A |
CERT: 3.086E-15 |
0.6629 |
 |

|
|
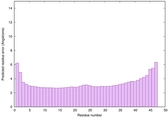
IntFOLD6 PPAS multi7 TS1 6h9mA2 3he5A 2oqqA 5al6A |
CERT: 3.757E-15 |
0.6621 |
 |

|
|
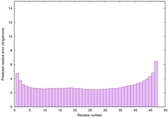
IntFOLD6 LOMETS multi3 TS1 4n6jA 5vr2A |
CERT: 4.052E-15 |
0.6617 |
 |

|
|
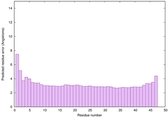
IntFOLD6 wPPAS multi8 TS1 5vr2A |
CERT: 4.39E-15 |
0.6614 |
 |

|
|
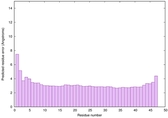
IntFOLD6 wPPAS multi6 TS1 5vr2A |
CERT: 4.39E-15 |
0.6614 |
 |

|
|
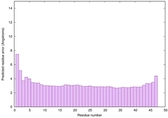
IntFOLD6 spk2 multi8 TS1 5vr2A |
CERT: 4.39E-15 |
0.6614 |
 |

|
|
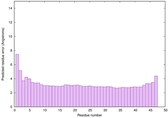
IntFOLD6 spk2 multi7 TS1 5vr2A |
CERT: 4.39E-15 |
0.6614 |
 |

|
|
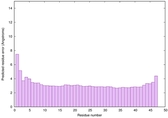
IntFOLD6 spk2 multi6 TS1 5vr2A |
CERT: 4.39E-15 |
0.6614 |
 |

|
|
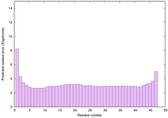
IntFOLD6 IntFOLDTS140 multi3 TS1 2x7aA 4pxjA 5al6A 5kb0A 4zqaA 2p4vA |
CERT: 5.308E-15 |
0.6605 |
 |

|
|
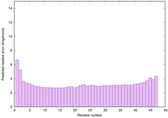
IntFOLD6 CNFsearch multi8 TS1 4wheA |
CERT: 5.751E-15 |
0.6602 |
 |

|
|
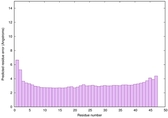
IntFOLD6 CNFsearch multi7 TS1 4wheA |
CERT: 5.751E-15 |
0.6602 |
 |

|
|
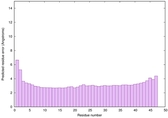
IntFOLD6 CNFsearch multi6 TS1 4wheA |
CERT: 5.751E-15 |
0.6602 |
 |

|
|
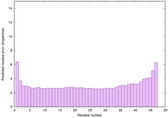
IntFOLD6 wdPPAS multi7 TS1 3he5A 2x7aA 4pxjA 5al6A 5kb0A |
CERT: 6.396E-15 |
0.6597 |
 |

|
|
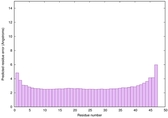
IntFOLD6 wMUSTER multi3 TS1 4n6jA 5vr2A 3he5A 2oqqA |
CERT: 8.254E-15 |
0.6586 |
 |

|
|
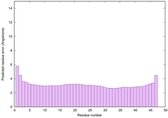
IntFOLD6 wdPPAS multi6 TS1 3he5A |
CERT: 8.903E-15 |
0.6583 |
 |

|
|
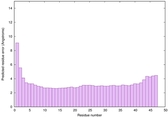
IntFOLD6 spk2 multi5 TS1 5vr2A 3he5B |
CERT: 1.02E-14 |
0.6577 |
 |

|
|
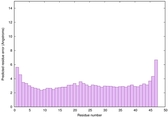
IntFOLD6 HHsearch multi1 TS1 4n6jA 3he5A |
CERT: 1.078E-14 |
0.6574 |
 |

|
|
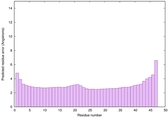
IntFOLD6 HHsearch multi7 TS1 4n6jA 4ywzA 4zqaA 2p4vA |
CERT: 1.198E-14 |
0.6570 |
 |

|
|
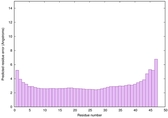
IntFOLD6 wPPAS multi4 TS1 4n6jA 5vr2A |
CERT: 1.726E-14 |
0.6553 |
 |

|
|
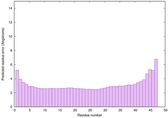
IntFOLD6 wPPAS multi1 TS1 4n6jA 5vr2A |
CERT: 1.726E-14 |
0.6553 |
 |

|
|
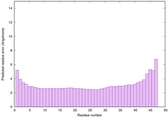
IntFOLD6 wMUSTER multi4 TS1 4n6jA 5vr2A |
CERT: 1.726E-14 |
0.6553 |
 |

|
|
IntFOLD6 wMUSTER multi1 TS1 4n6jA 5vr2A |
CERT: 1.726E-14 |
0.6553 |
 |

|
|
IntFOLD6 SPARKSX multi1 TS1 4n6jA 4hb1A |
CERT: 2.128E-14 |
0.6544 |
 |

|
|
IntFOLD6 wMUSTER multi8 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
 |

|
|
IntFOLD6 wMUSTER multi6 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
 |

|
|
IntFOLD6 PPAS multi8 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
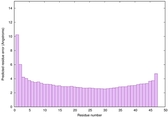 |

|
|
IntFOLD6 PPAS multi6 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
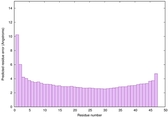 |

|
|
IntFOLD6 MUSTER multi8 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
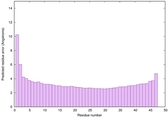 |

|
|
IntFOLD6 MUSTER multi6 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
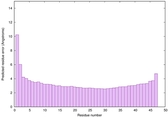 |

|
|
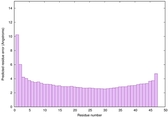
IntFOLD6 dPPAS multi8 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
 |

|
|
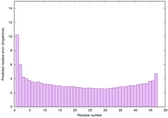
IntFOLD6 dPPAS multi6 TS1 6h9mA2 |
CERT: 2.211E-14 |
0.6543 |
 |

|
|
IntFOLD6 spk2 multi3 TS1 4n6jA 3he5B 5vr2A |
CERT: 2.352E-14 |
0.6540 |
 |

|
|
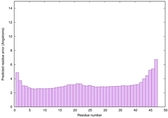
IntFOLD6 HHsearch multi8 TS1 4n6jA 4ywzA |
CERT: 2.722E-14 |
0.6533 |
 |

|
|
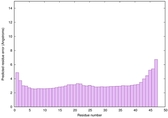
IntFOLD6 HHsearch multi4 TS1 4n6jA 4ywzA |
CERT: 2.722E-14 |
0.6533 |
 |

|
|
IntFOLD6 wPPAS multi7 TS1 5vr2A 4pxjA |
CERT: 2.755E-14 |
0.6533 |
 |

|
|
IntFOLD6 wPPAS multi5 TS1 5vr2A 5f5sB |
CERT: 3.366E-14 |
0.6524 |
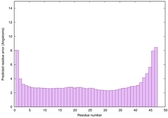 |

|
|
IntFOLD6 MUSTER multi3 TS1 4n6jA 5vr2A 3he5A 2oqqA |
CERT: 4.595E-14 |
0.6510 |
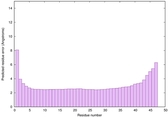 |

|
|
IntFOLD6 HHsearch multi5 TS1 4n6jA 3b8mA |
CERT: 4.897E-14 |
0.6508 |
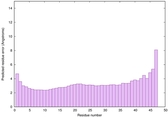 |

|
|
IntFOLD6 dPPAS2 multi3 TS1 4n6jA 5vr2A 4pxjA |
CERT: 5.126E-14 |
0.6506 |
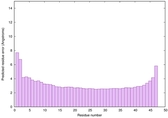 |

|
|
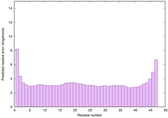
IntFOLD6 dPPAS2 multi6 TS1 4pkyB |
CERT: 6.075E-14 |
0.6498 |
 |

|
|
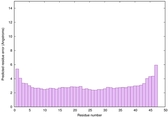
IntFOLD6 SPARKSX multi8 TS1 4n6jA 5vnyA |
CERT: 6.253E-14 |
0.6497 |
 |

|
|
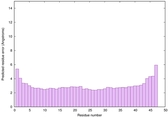
IntFOLD6 SPARKSX multi5 TS1 4n6jA 5vnyA |
CERT: 6.253E-14 |
0.6497 |
 |

|
|
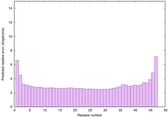
IntFOLD6 wdPPAS multi4 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
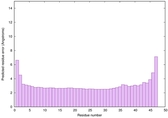
IntFOLD6 wdPPAS multi1 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
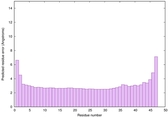
IntFOLD6 PPAS multi4 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
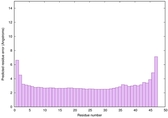
IntFOLD6 PPAS multi1 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
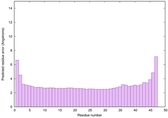
IntFOLD6 MUSTER multi4 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
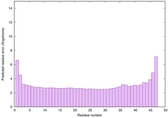
IntFOLD6 MUSTER multi1 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
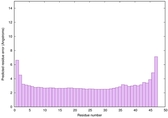
IntFOLD6 LOMETS multi4 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
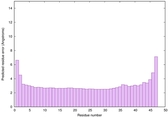
IntFOLD6 dPPAS multi4 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
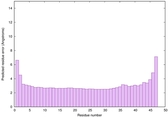
IntFOLD6 dPPAS multi1 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
 |

|
|
IntFOLD6 dPPAS2 multi4 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
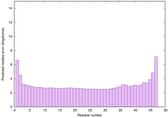 |

|
|
IntFOLD6 dPPAS2 multi1 TS1 4n6jA 5vr2A |
CERT: 5.646E-13 |
0.6400 |
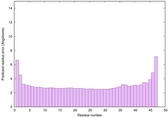 |

|
|
IntFOLD6 PPAS multi3 TS1 4n6jA 5vr2A 2oqqA 4pkyB 3he5A 5al6A |
CERT: 7.147E-13 |
0.6389 |
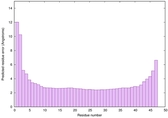 |

|
|
IntFOLD6 spk2 multi4 TS1 4n6jA 3he5B |
CERT: 1.563E-11 |
0.6254 |
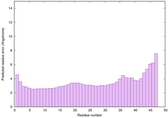 |

|
|
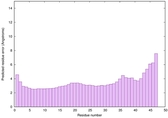
IntFOLD6 spk2 multi1 TS1 4n6jA 3he5B |
CERT: 1.563E-11 |
0.6254 |
 |

|
|
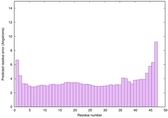
IntFOLD6 wdPPAS multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
 |

|
|
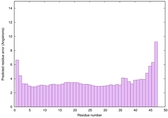
IntFOLD6 PPAS multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
 |

|
|
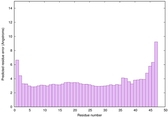
IntFOLD6 MUSTER multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
 |

|
|
IntFOLD6 LOMETS multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
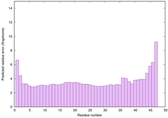 |

|
|
IntFOLD6 LOMETS multi1 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
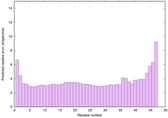 |

|
|
IntFOLD6 Env-PPAS multi7 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
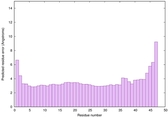 |

|
|
IntFOLD6 Env-PPAS multi6 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
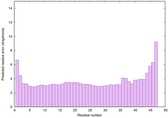 |

|
|
IntFOLD6 Env-PPAS multi3 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
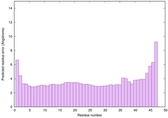 |

|
|
IntFOLD6 Env-PPAS multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
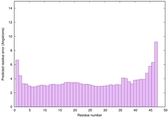 |

|
|
IntFOLD6 dPPAS multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
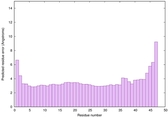 |

|
|
IntFOLD6 dPPAS2 multi2 TS1 4n6jA |
CERT: 2.366E-11 |
0.6235 |
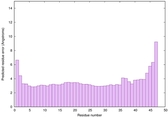 |

|
|
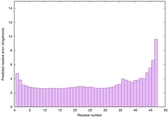
IntFOLD6 sp3 multi7 TS1 4n6jA |
CERT: 8.21E-11 |
0.6181 |
 |

|
|
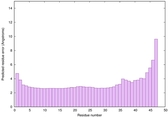
IntFOLD6 sp3 multi6 TS1 4n6jA |
CERT: 8.21E-11 |
0.6181 |
 |

|
|
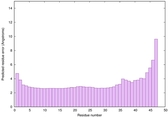
IntFOLD6 sp3 multi3 TS1 4n6jA |
CERT: 8.21E-11 |
0.6181 |
 |

|
|
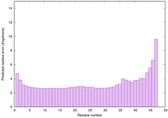
IntFOLD6 sp3 multi2 TS1 4n6jA |
CERT: 8.21E-11 |
0.6181 |
 |

|
|
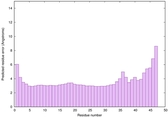
IntFOLD6 wPPAS multi2 TS1 4n6jA |
CERT: 8.6E-11 |
0.6178 |
 |

|
|
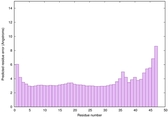
IntFOLD6 wMUSTER multi2 TS1 4n6jA |
CERT: 8.6E-11 |
0.6178 |
 |

|
|
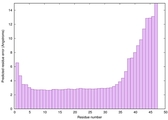
IntFOLD6 SPARKSX multi4 TS1 4n6jA 2gomA |
CERT: 7.307E-8 |
0.5881 |
 |

|
|
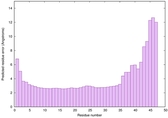
IntFOLD6 spk2 multi2 TS1 4n6jA |
CERT: 2.57E-7 |
0.5826 |
 |

|
|
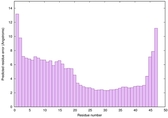
IntFOLD6 sp3 multi8 TS1 4n6jA 1a32 |
CERT: 3.205E-7 |
0.5816 |
 |

|
|
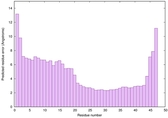
IntFOLD6 sp3 multi5 TS1 4n6jA 1a32 |
CERT: 3.205E-7 |
0.5816 |
 |

|
|
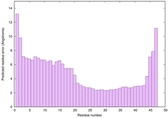
IntFOLD6 sp3 multi4 TS1 4n6jA 1a32 |
CERT: 3.205E-7 |
0.5816 |
 |

|
|
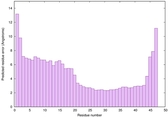
IntFOLD6 sp3 multi1 TS1 4n6jA 1a32 |
CERT: 3.205E-7 |
0.5816 |
 |

|
|
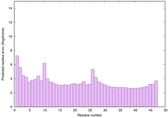
IntFOLD6 DMPf TS1 |
CERT: 6.055E-7 |
0.5788 |
 |

|
|
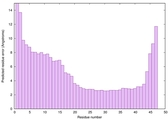
IntFOLD6 Env-PPAS multi8 TS1 4n6jA 3bbnO |
CERT: 5.608E-5 |
0.5589 |
 |

|
|
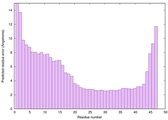
IntFOLD6 Env-PPAS multi5 TS1 4n6jA 3bbnO |
CERT: 5.608E-5 |
0.5589 |
 |

|
|
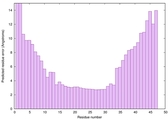
IntFOLD6 Env-PPAS multi4 TS1 4n6jA 5oeuA2 |
HIGH: 1.414E-3 |
0.5447 |
 |
|
|
IntFOLD6 Env-PPAS multi1 TS1 4n6jA 5oeuA2 |
HIGH: 1.414E-3 |
0.5447 |
 |
|