The IntFOLD Server Results Page (Version 5.0)
Please cite the relevant papers:McGuffin, L.J., Adiyaman, R., Maghrabi, A.H.A., Shuid, A.N., Brackenridge, D.A., Nealon, J.O. & Philomina, L.S. (2019) IntFOLD: an integrated web resource for high performance protein structure and function prediction. Nucleic Acids Research, 47, W408-W413. DOI PubMed
McGuffin, L.J., Shuid, A.M., Kempster, R., Maghrabi, A.H.A., Nealon J.O., Salehe, B.R., Atkins, J.D. & Roche, D.B. (2017) Accurate Template Based Modelling in CASP12 using the IntFOLD4-TS, ModFOLD6 and ReFOLD methods. Proteins: Structure, Function, and Bioinformatics, 86 Suppl 1, 335-344, doi: 10.1002/prot.25360. PubMed
Links to graphical output:
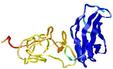
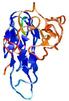
- Top 5 3D models
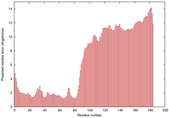
- Disorder prediction
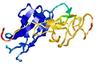
- Domain boundary prediction
- Binding site prediction
- Full model quality assessment results
Download machine readable results in CASP format:
- TS (Tertiary Structure Prediction)
- DR (Disorder Prediction)
- DP (Domain Prediction)
- FN (Binding Site Prediction)
- QA (Model Quality Prediction)
- Make a compressed file containing all generated 3D models (models are in PDB format with predicted per-residue errors in the B-factor column)
Results will be available for 21 days after job completion (subject to server capacity)
| Tertiary Structure Predictions: Top 5 multi-template 3D models for T0964 | Help | ||||
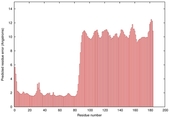
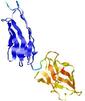
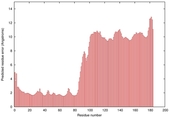
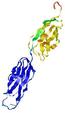
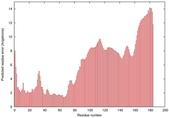
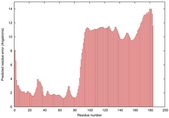
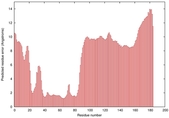
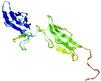
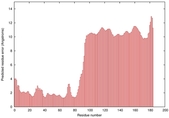
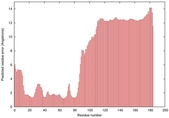
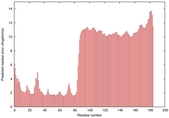
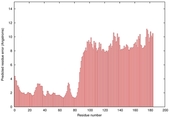
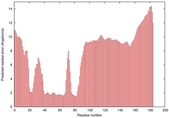
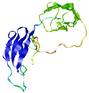
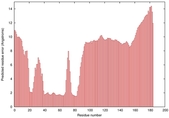
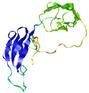
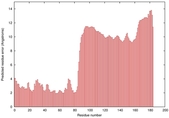
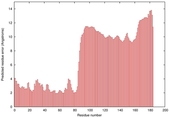
| Model ID and PDBsum links for all templates used | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | 3D map of model quality |
|
IntFOLD5 wdPPAS multi4 TS1 2l04A 5ngjA2 5t7aA |
CERT: 6.497E-5 |
0.4222 |
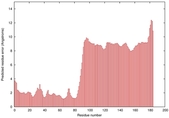 |

|
|
IntFOLD5 wdPPAS multi6 TS1 5ngjA2 5t7aA |
CERT: 8.18E-5 |
0.4190 |
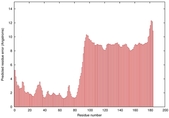 |
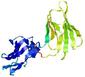
|
|
IntFOLD5 wMUSTER multi8 TS1 5ngjA 5ldyA |
CERT: 1.019E-4 |
0.4160 |
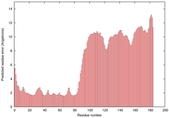 |

|
|
IntFOLD5 HHpred TS2 5ftxA 5h9xA 2l04A 2mh4A 2mogA |
CERT: 1.167E-4 |
0.4142 |
 |
|
|
IntFOLD5 dPPAS2 multi8 TS1 2l04A 5ngjA2 5ldyA |
CERT: 1.417E-4 |
0.4116 |
 |
|
McGuffin, L. J. (2008) Intrinsic disorder prediction from the analysis of multiple protein fold recognition models. Bioinformatics, 24, 1789-1804. PubMed
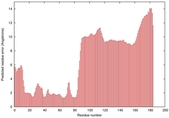
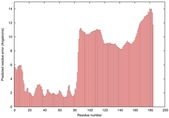
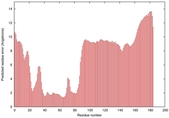
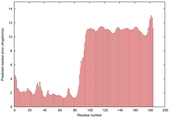
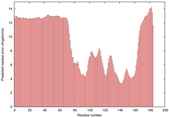
| Disorder prediction for T0964 | Help | |||
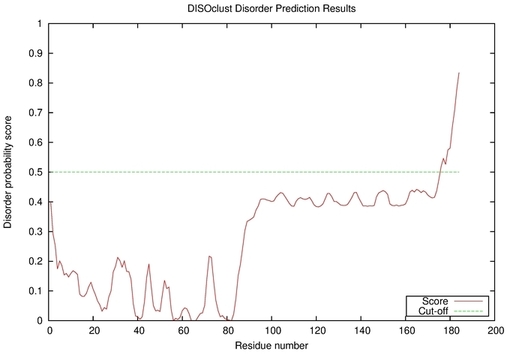
| |||
| Click image to download plot in PostScript format. |
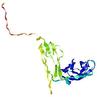
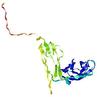
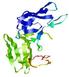
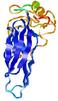
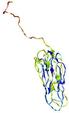
| Domain boundary prediction for T0964 | Help | |||
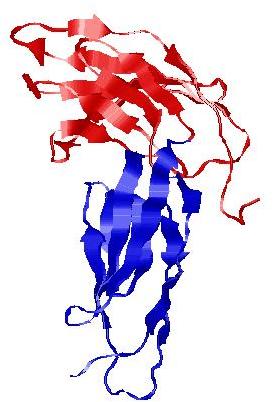
| |||
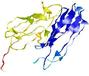
| Click image to download model or view prediction using Jmol. The model above is coloured according to the predicted domains. |
Roche, D. B., Tetchner, S. J. & McGuffin, L. J. (2011) FunFOLD: an improved automated method for the prediction of ligand binding residues using 3D models of proteins. BMC Bioinformatics, 12, 160. PubMed
| Ligand binding residues prediction for T0964 | Help | |||
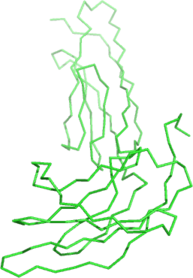
| |||
| No binding residues predicted. |
Maghrabi, A.H.A. & McGuffin L.J. (2017) ModFOLD6: an accurate web server for the global and local quality estimation of 3D models of proteins. Nucleic Acids Res., 45, W416-W421, doi: 10.1093/nar/gkx332. PubMed
| Full model quality assessment results for T0964 | Help | ||||
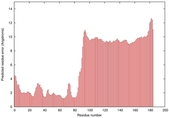
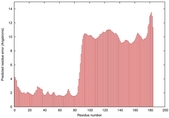
| Model ID and PDBsum links for all templates used | Confidence and P-value | Global model quality score | Local model quality plot (click images to download plots) | 3D map of model quality |
|
IntFOLD5 wdPPAS multi4 TS1 2l04A 5ngjA2 5t7aA |
CERT: 6.497E-5 |
0.4222 |
 |

|
|
IntFOLD5 wdPPAS multi6 TS1 5ngjA2 5t7aA |
CERT: 8.18E-5 |
0.4190 |
 |

|
|
IntFOLD5 wMUSTER multi8 TS1 5ngjA 5ldyA |
CERT: 1.019E-4 |
0.4160 |
 |

|
|
IntFOLD5 HHpred TS2 5ftxA 5h9xA 2l04A 2mh4A 2mogA |
CERT: 1.167E-4 |
0.4142 |
 |

|
|
IntFOLD5 dPPAS2 multi8 TS1 2l04A 5ngjA2 5ldyA |
CERT: 1.417E-4 |
0.4116 |
 |

|
|
IntFOLD5 dPPAS multi8 TS1 5ngjA2 2l04A 5ldyA |
CERT: 1.56E-4 |
0.4103 |
 |

|
|
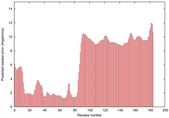
IntFOLD5 CNFsearch multi4 TS1 5ngjA 4uidA 4uj7A 1cwvA |
CERT: 3.156E-4 |
0.4010 |
 |

|
|
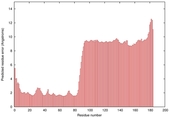
IntFOLD5 MUSTER multi4 TS1 2l04A 5ngjA2 3pddA |
CERT: 3.197E-4 |
0.4008 |
 |

|
|
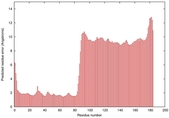
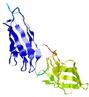
IntFOLD5 HHpred TS3 5ftxA 5h9xA 2l04A 2mh4A 2mogA |
CERT: 3.394E-4 |
0.4001 |
 |
|
|
IntFOLD5 CNFsearch multi1 TS1 5ngjA 4uidA |
CERT: 4.186E-4 |
0.3973 |
 |

|
|
IntFOLD5 wdPPAS multi2 TS1 2l04A 5t7aA |
CERT: 6.482E-4 |
0.3917 |
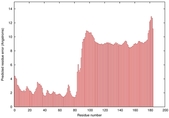 |

|
|
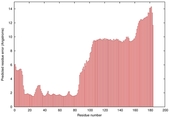
IntFOLD5 PPAS multi4 TS1 2l04A 5ngjA2 3pddA |
CERT: 8.805E-4 |
0.3878 |
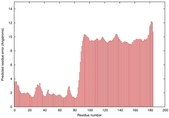 |
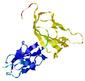
|
|
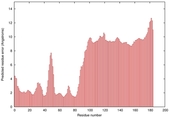
IntFOLD5 spk2 multi8 TS1 2l04A 3pddA |
CERT: 9.86E-4 |
0.3864 |
 |

|
|
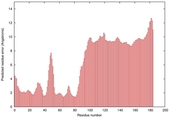
IntFOLD5 spk2 multi5 TS1 2l04A 3pddA |
CERT: 9.86E-4 |
0.3864 |
 |

|
|
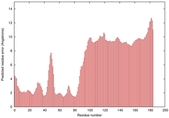
IntFOLD5 spk2 multi4 TS1 2l04A 3pddA |
CERT: 9.86E-4 |
0.3864 |
 |

|
|
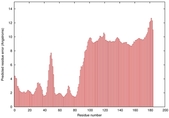
IntFOLD5 spk2 multi1 TS1 2l04A 3pddA |
CERT: 9.86E-4 |
0.3864 |
 |

|
|
IntFOLD5 PPAS multi8 TS1 5ngjA2 2l04A 2zqkB 5ldyA |
HIGH: 1.009E-3 |
0.3861 |
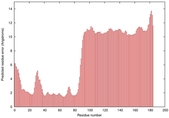 |

|
|
IntFOLD5 sp3 multi8 TS1 2l04A 3pddA |
HIGH: 1.047E-3 |
0.3857 |
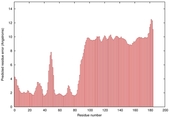 |

|
|
IntFOLD5 sp3 multi5 TS1 2l04A 3pddA |
HIGH: 1.047E-3 |
0.3857 |
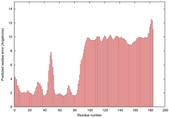 |

|
|
IntFOLD5 wMUSTER multi3 TS1 5ftxA 5ngjA 2l04A |
HIGH: 1.088E-3 |
0.3852 |
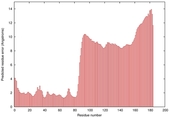 |
|
|
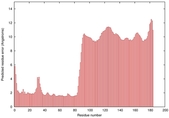
IntFOLD5 CNFsearch multi6 TS1 4uj7A |
HIGH: 1.18E-3 |
0.3842 |
 |

|
|
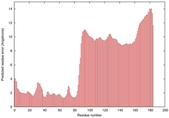
IntFOLD5 COMA multi5 TS1 2l04A 6qx4A |
HIGH: 1.456E-3 |
0.3816 |
 |

|
|
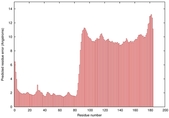
IntFOLD5 HHpred TS1 5ftxA 5h9xA 2l04A 2mh4A 2mogA |
HIGH: 1.495E-3 |
0.3812 |
 |
|
|
IntFOLD5 dPPAS multi4 TS1 2l04A 5ngjA2 3pddA |
HIGH: 1.71E-3 |
0.3796 |
 |
|
|
IntFOLD5 CNFsearch multi7 TS1 4uj7A 2l04A |
HIGH: 1.718E-3 |
0.3795 |
 |

|
|
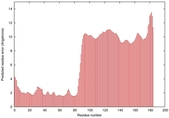
IntFOLD5 CNFsearch multi5 TS1 4uj7A 2l04A |
HIGH: 1.718E-3 |
0.3795 |
 |

|
|
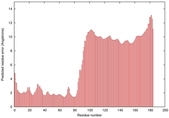
IntFOLD5 IT5 TS3 |
HIGH: 3.015E-3 |
0.3726 |
 |
|
|
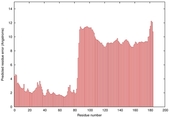
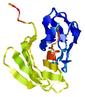
IntFOLD5 wdPPAS multi8 TS1 5ngjA2 2l04A 4hu8A1 5t7aA |
HIGH: 3.102E-3 |
0.3723 |
 |
|
|
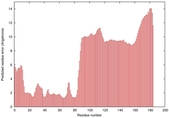
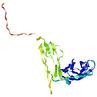
IntFOLD5 HHsearch multi8 TS1 2l04A 5ngjA |
HIGH: 9.931E-3 |
0.3585 |
 |
|
|
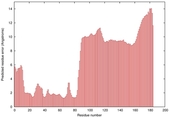
IntFOLD5 HHsearch multi6 TS1 2l04A 5ngjA |
HIGH: 9.931E-3 |
0.3585 |
 |
|
|
IntFOLD5 HHsearch multi5 TS1 2l04A 5ngjA |
HIGH: 9.931E-3 |
0.3585 |
 |
|
|
IntFOLD5 HHsearch multi4 TS1 4uidA 5ngjA 2l04A |
MEDIUM: 1.366E-2 |
0.3548 |
 |

|
|
IntFOLD5 HHsearch multi2 TS1 4uidA 2l04A |
MEDIUM: 1.574E-2 |
0.3531 |
 |

|
|
IntFOLD5 COMA multi8 TS1 2l04A 6qx4A 2zqkA 1f00I |
MEDIUM: 1.579E-2 |
0.3531 |
 |

|
|
IntFOLD5 SPARKSX multi1 TS1 6n1aA 4uidA |
MEDIUM: 2.019E-2 |
0.3503 |
 |
|
|
IntFOLD5 IT5 TS1 |
MEDIUM: 2.051E-2 |
0.3501 |
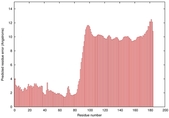 |

|
|
IntFOLD5 dPPAS multi6 TS1 5ngjA2 5ldyA2 |
MEDIUM: 2.112E-2 |
0.3498 |
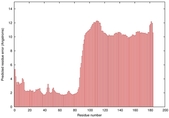 |

|
|
IntFOLD5 wPPAS multi4 TS1 2l04A 5ngjA2 4hu8A1 2zqkB 3pddA |
MEDIUM: 2.248E-2 |
0.3490 |
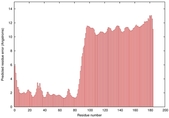 |
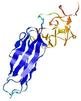
|
|
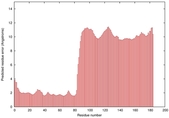
IntFOLD5 sp3 multi4 TS1 2l04A 6ndtB 3pddA |
MEDIUM: 2.639E-2 |
0.3469 |
 |
|
|
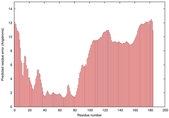
IntFOLD5 IT5 TS2 |
MEDIUM: 2.792E-2 |
0.3462 |
 |

|
|
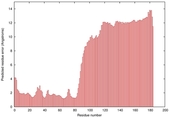
IntFOLD5 PPAS multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 3.098E-2 |
0.3447 |
 |

|
|
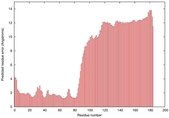
IntFOLD5 Env-PPAS multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 3.098E-2 |
0.3447 |
 |

|
|
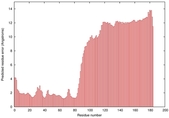
IntFOLD5 dPPAS2 multi5 TS1 2l04A 5ngjA2 |
MEDIUM: 3.098E-2 |
0.3447 |
 |

|
|
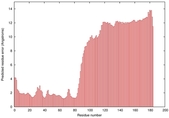
IntFOLD5 dPPAS2 multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 3.098E-2 |
0.3447 |
 |

|
|
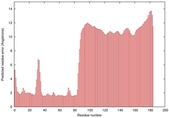
IntFOLD5 CNFsearch multi8 TS1 4uj7A 1f00I |
MEDIUM: 3.203E-2 |
0.3442 |
 |
|
|
IntFOLD5 SPARKSX multi5 TS1 2l04A 5ngjA |
MEDIUM: 3.252E-2 |
0.3440 |
 |

|
|
IntFOLD5 Env-PPAS multi8 TS1 5ngjA2 2l04A 2zqkB 1cwvA4 |
MEDIUM: 3.382E-2 |
0.3434 |
 |
|
|
IntFOLD5 wdPPAS multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 3.413E-2 |
0.3432 |
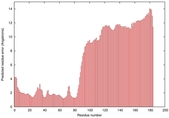 |

|
|
IntFOLD5 dPPAS multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 3.413E-2 |
0.3432 |
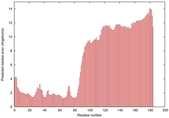 |

|
|
IntFOLD5 wMUSTER multi4 TS1 5ftxA |
MEDIUM: 3.697E-2 |
0.3419 |
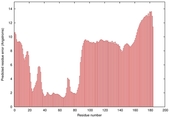 |
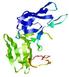
|
|
IntFOLD5 wMUSTER multi2 TS1 5ftxA |
MEDIUM: 3.697E-2 |
0.3419 |
 |
|
|
IntFOLD5 Env-PPAS multi4 TS1 2l04A 5ngjA2 2zqkB 1cwvA4 |
MEDIUM: 3.724E-2 |
0.3418 |
 |
|
|
IntFOLD5 wPPAS multi7 TS1 5ngjA2 2l04A |
MEDIUM: 4.202E-2 |
0.3398 |
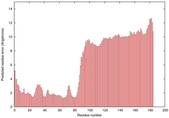 |

|
|
IntFOLD5 wPPAS multi5 TS1 5ngjA2 2l04A |
MEDIUM: 4.202E-2 |
0.3398 |
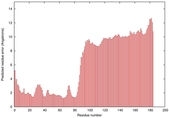 |

|
|
IntFOLD5 MUSTER multi6 TS1 5ngjA2 |
MEDIUM: 4.213E-2 |
0.3397 |
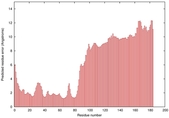 |

|
|
IntFOLD5 wdPPAS multi7 TS1 5ngjA2 2l04A |
MEDIUM: 4.291E-2 |
0.3394 |
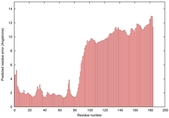 |

|
|
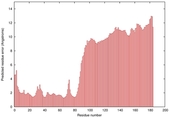
IntFOLD5 wdPPAS multi5 TS1 5ngjA2 2l04A |
MEDIUM: 4.291E-2 |
0.3394 |
 |

|
|
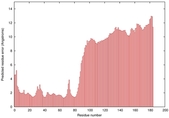
IntFOLD5 dPPAS multi7 TS1 5ngjA2 2l04A |
MEDIUM: 4.291E-2 |
0.3394 |
 |

|
|
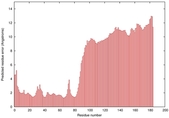
IntFOLD5 dPPAS multi5 TS1 5ngjA2 2l04A |
MEDIUM: 4.291E-2 |
0.3394 |
 |

|
|
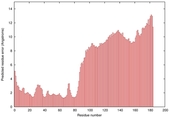
IntFOLD5 wMUSTER multi6 TS1 5ngjA |
MEDIUM: 4.358E-2 |
0.3391 |
 |

|
|
IntFOLD5 MUSTER multi7 TS1 5ngjA2 2l04A |
MEDIUM: 4.376E-2 |
0.3390 |
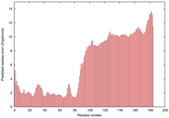 |

|
|
IntFOLD5 MUSTER multi5 TS1 5ngjA2 2l04A |
MEDIUM: 4.376E-2 |
0.3390 |
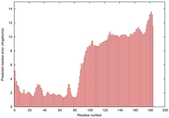 |

|
|
IntFOLD5 wMUSTER multi7 TS1 5ngjA 2l04A |
MEDIUM: 4.41E-2 |
0.3389 |
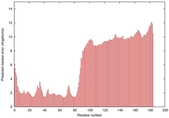 |

|
|
IntFOLD5 wPPAS multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 4.418E-2 |
0.3388 |
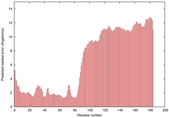 |

|
|
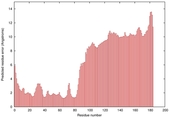
IntFOLD5 PPAS multi6 TS1 5ngjA2 |
MEDIUM: 4.484E-2 |
0.3386 |
 |
|
|
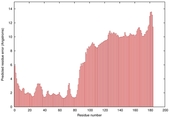
IntFOLD5 Env-PPAS multi6 TS1 5ngjA2 |
MEDIUM: 4.484E-2 |
0.3386 |
 |
|
|
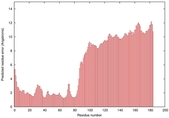
IntFOLD5 wMUSTER multi5 TS1 5ngjA 5ngjA2 |
MEDIUM: 4.591E-2 |
0.3381 |
 |

|
|
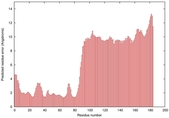
IntFOLD5 PPAS multi7 TS1 5ngjA2 2l04A |
MEDIUM: 4.655E-2 |
0.3379 |
 |

|
|
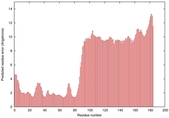
IntFOLD5 PPAS multi5 TS1 5ngjA2 2l04A |
MEDIUM: 4.655E-2 |
0.3379 |
 |

|
|
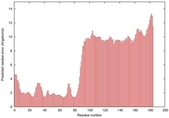
IntFOLD5 Env-PPAS multi7 TS1 5ngjA2 2l04A |
MEDIUM: 4.655E-2 |
0.3379 |
 |

|
|
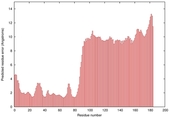
IntFOLD5 Env-PPAS multi5 TS1 5ngjA2 2l04A |
MEDIUM: 4.655E-2 |
0.3379 |
 |

|
|
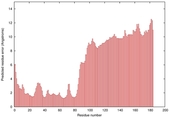
IntFOLD5 wPPAS multi6 TS1 5ngjA2 |
MEDIUM: 4.738E-2 |
0.3375 |
 |

|
|
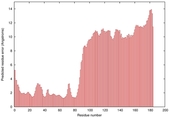
IntFOLD5 MUSTER multi1 TS1 2l04A 5ngjA2 |
MEDIUM: 4.759E-2 |
0.3374 |
 |

|
|
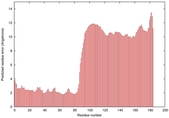
IntFOLD5 dPPAS multi2 TS1 2l04A 5ldyA2 |
LOW: 5.597E-2 |
0.3341 |
 |
|
|
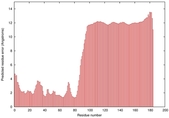
IntFOLD5 wPPAS multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
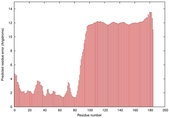
IntFOLD5 wPPAS multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
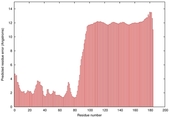
IntFOLD5 wdPPAS multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
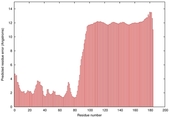
IntFOLD5 spk2 multi7 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
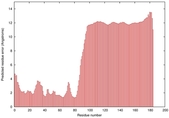
IntFOLD5 spk2 multi6 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
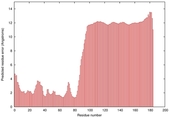
IntFOLD5 spk2 multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
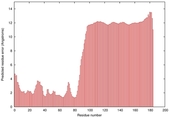
IntFOLD5 spk2 multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
IntFOLD5 SPARKSX multi7 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
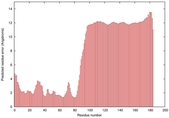 |

|
|
IntFOLD5 SPARKSX multi6 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
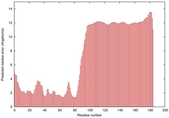 |

|
|
IntFOLD5 sp3 multi7 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
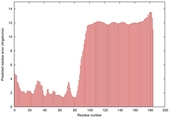 |

|
|
IntFOLD5 sp3 multi6 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
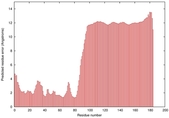 |

|
|
IntFOLD5 sp3 multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
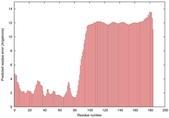
IntFOLD5 sp3 multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
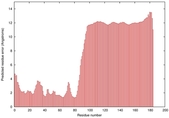
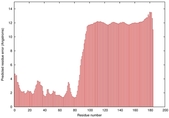
IntFOLD5 PPAS multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
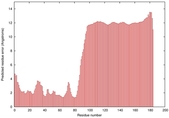
IntFOLD5 PPAS multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
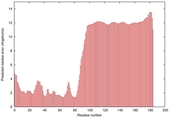
IntFOLD5 MUSTER multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
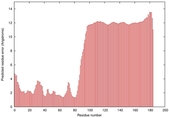
IntFOLD5 MUSTER multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
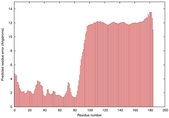
IntFOLD5 LOMETS multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
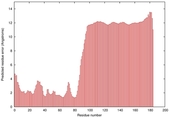
IntFOLD5 LOMETS multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
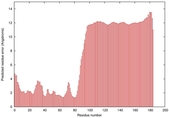
IntFOLD5 IntFOLDTS60 multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
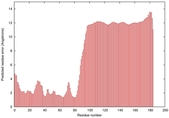
IntFOLD5 IntFOLDTS60 multi1 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
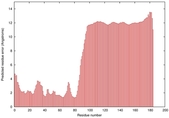
IntFOLD5 IntFOLDTS140 multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
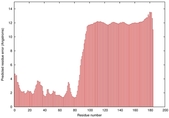
IntFOLD5 IntFOLDTS140 multi1 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
IntFOLD5 Env-PPAS multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
IntFOLD5 Env-PPAS multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
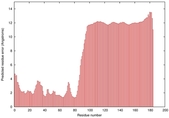
IntFOLD5 dPPAS multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
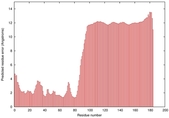
IntFOLD5 dPPAS2 multi7 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
 |

|
|
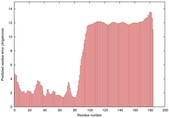
IntFOLD5 dPPAS2 multi6 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
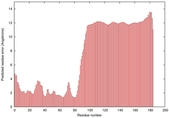 |

|
|
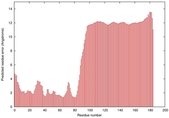
IntFOLD5 dPPAS2 multi3 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
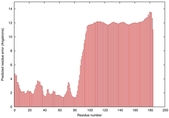 |

|
|
IntFOLD5 dPPAS2 multi2 TS1 2l04A |
LOW: 5.698E-2 |
0.3337 |
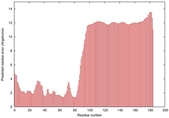 |

|
|
IntFOLD5 CNFsearch multi3 TS1 5ngjA 2l04A |
LOW: 5.927E-2 |
0.3328 |
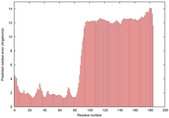 |

|
|
IntFOLD5 wMUSTER multi1 TS1 5ftxA 5ngjA |
LOW: 6.026E-2 |
0.3324 |
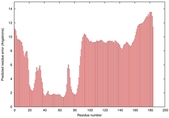 |
|
|
IntFOLD5 COMA multi3 TS1 4uidA 2l04A |
LOW: 6.136E-2 |
0.3320 |
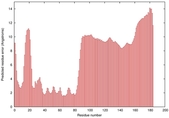 |

|
|
IntFOLD5 COMA multi1 TS1 4uidA 2l04A |
LOW: 6.136E-2 |
0.3320 |
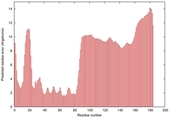 |

|
|
IntFOLD5 CNFsearch multi2 TS1 5ngjA |
LOW: 6.282E-2 |
0.3314 |
 |

|
|
IntFOLD5 dPPAS2 multi4 TS1 2l04A 5ngjA2 4uicA3 5ldyA |
LOW: 6.442E-2 |
0.3308 |
 |
|
|
IntFOLD5 COMA multi7 TS1 2l04A |
LOW: 7.248E-2 |
0.3278 |
 |

|
|
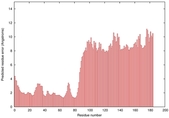
IntFOLD5 COMA multi6 TS1 2l04A |
LOW: 7.248E-2 |
0.3278 |
 |

|
|
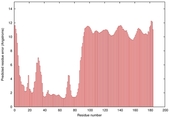
IntFOLD5 MUSTER multi8 TS1 5ngjA2 2zqkB |
LOW: 7.831E-2 |
0.3256 |
 |

|
|
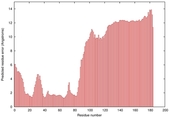
IntFOLD5 HHsearch multi3 TS1 4uidA 5ngjA |
LOW: 8.133E-2 |
0.3245 |
 |

|
|
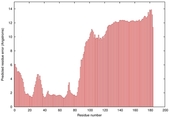
IntFOLD5 HHsearch multi1 TS1 4uidA 5ngjA |
LOW: 8.133E-2 |
0.3245 |
 |

|
|
IntFOLD5 LOMETS multi4 TS1 2l04A 5ftxA |
LOW: 8.249E-2 |
0.3241 |
 |
|
|
IntFOLD5 LOMETS multi1 TS1 2l04A 5ftxA |
LOW: 8.249E-2 |
0.3241 |
 |
|
|
IntFOLD5 COMA multi2 TS1 4uidA |
LOW: 8.319E-2 |
0.3239 |
 |

|
|
IntFOLD5 wPPAS multi8 TS1 5ngjA2 2zqkB 3pddA |
UNCERT: 1.097E-1 |
0.3152 |
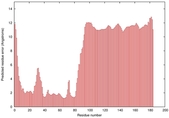 |

|
|
IntFOLD5 COMA multi4 TS1 4uidA 5ngjA 1f00I |
UNCERT: 2.025E-1 |
0.2948 |
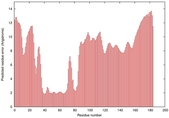 |
|
|
IntFOLD5 sp3 multi1 TS1 2l04A 6ndtB |
UNCERT: 2.684E-1 |
0.2852 |
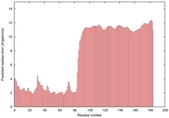 |
|
|
IntFOLD5 IntFOLDTS60 multi2 TS1 2l04A |
UNCERT: 4.969E-1 |
0.2607 |
 |
|
|
IntFOLD5 IntFOLDTS140 multi2 TS1 2l04A |
UNCERT: 4.969E-1 |
0.2607 |
 |
|
|
IntFOLD5 HHsearch multi7 TS1 2l04A |
UNCERT: 8.037E-1 |
0.2255 |
 |

|